Reverse transcriptase and intron number evolution
Introduction
Introns are universal in eukaryotic genomes and play important roles in transcriptional regulation (1-3), mRNA export to the cytoplasm (4), nonsense-mediated decay as both a regulatory and a splicing quality control mechanism (5-7), R-loop avoidance (8), alternative splicing (9-11), chromatin structure (12-17), and evolution by exon-shuffling (18-21). Fungal genomes mostly range from 10 to 90 million base pairs (22) in size and vary widely in intron densities. In the low end, the model yeast Saccharomyces cerevisiae has 283 intron-containing genes, and only about 20% of intron deletions caused minor phenotypes under different growth conditions, which led to the view that many introns can be phase out without blocking cell growth (23). On the high end, Basidiomycota yeast Sporobolomyces roseus has over seven coding exons per gene (EPG) (http://www.jgi.doe/Sroseus). Other fungal genomes have intron densities in between the two extremes (24-26). Therefore, fungal genomes are well suited for studying intron evolution.
Introns shared in diverse eukaryotes are inherited from a common intron-rich ancestor whose descendants have gone through mainly intron loss of varying degree (27-36). Investigations of insertion and deletion of introns from orthologous gene sets (32,37-41) have led to a consensual view that intron loss is from one to several orders of magnitude more frequent than intron gain (25,31,36,37,42-48). The gain and loss rates are lineage-specific (49). Intron loss has been attributed to reverse transcriptase (RT) (43,50,51). A cDNA generated by RT from an mRNA undergoes homologous recombination with its parent gene and leads to intron loss (43,48) with a 3'-bias (52) that is supported by enzyme kinetics (53,54). Self-priming (52) [as opposed to external priming (55,56) and 5'-self-priming (57)], together with high processivity of RT as exemplified by the human L1 non-LTR type (54), favors more intron loss in the middle of genes. Lack of 3' introns could alternatively be explained by nonsense-mediated mRNA decay (NMD) where introns after a stop codon could cause rapid degradation of mRNA (5,58).
The rate of intron gain and loss have been assessed with parsimony (59) and likelihood (32,37) methods. These mature methods (27,39,60,61) treat intron presence/absence as two character states in the context of multiple protein sequence alignment and are instrumental to understanding intron evolution. For example, they tell us the intron density of last eukaryotic common ancestor (LECA) with an upward trend in chronology (25,28,31,37,40). However, there might be limitations as discussed in the context of evolutionary biology (33). Moreover, the sampling space may be biased for these methods because they pick introns from highly conserved gap-free regions of multiple sequence alignments. Armed with two observations: conservation of the mode of exon length distribution and relative intron location (RIL) as a tracer for RT-mediated intron loss, we were able to look into the intron states of the ancestral eukaryotes from different angles. The intron density of the ancestral genomes (7.7 or 16 EPG) determined with our intuitive method is significantly higher than the most recent estimates of 6.7 EPG and 6.4 EPG (40) for ancestors of fungi and eukaryotes, respectively.
Methods
Data
We used 16 complete fungal genomes (Table 1), 13 of which were sequenced and annotated by JGI (http://jgi.doe.gov/fungi). Ustilago maydis, Cryptococcus neoformans, and Coprinus cinereus (also named Coprinopsis cinerea) were from the Broad Institute (http://www.broadinstitute.org). Gene models from JGI-annotated genomes were taken from the GeneCatalog track that contained the best representative gene models. Varying fractions of the GeneCatalog were manually curated by external users. For clarity, we used the JGI unique database identifier (which consists of the first three letters from the genus name and two letters from the species name followed by a version number) in figures and tables.
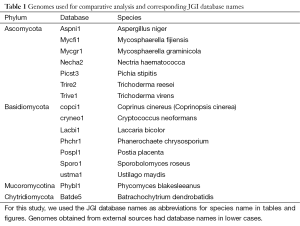
Full table
Clusters of proteins
We first did all-against-all Blastp (http://blast.ncbi.nlm.nih.gov/Blast.cgi) of protein sequences. Then we selected the top 10% of the blast hits for each protein and carried out Smith-Waterman alignments (62) on the identified pairs. After removing poor alignments (usually caused by low complexity regions), we picked the alignments with the best alignment score for each query sequence against each target database. We retained only one sequence if multiple sequences shared more than 98.5% sequence identity and more than 90% coverage from the same genome. The pairwise orthologous relationship was used to generate protein clusters with the single-linkage algorithm.
The clusters were divided into four categories: “all” (conserved in all species), “between” (conserved between Phyla), “phylum” (conserved within Phylum), and “species” (species-specific). For operational purposes, the two species from Mucoromycotina and Chytridiomycota were considered as one Phylum because in the unrooted phylogenetic tree of the 16 genomes, Phycomyces blakesleeanus (Phybl1) and Batrachochytrium dendrobatidis (Batde5) belonged to the same clade; this was supported by independent methods (63). The two species Trichoderma reesei and Trichoderma virens are closely related, so we will consider clusters containing proteins only from the above two species as species-specific.
Phylogenetic tree construction
We picked 288 protein clusters that contained exactly one member from each of the 16 genomes. Then we did all pairwise Smith-Waterman alignments, the average non-gapped sequence identity was computed for 120 genome-to-genome pairs. We used the Grishin distance calculation formula: q=ln(1+2d)/[2d], where q is the fraction of identical residues between two aligned sequences, and d is the amino acid substitution per site (64). The tree was constructed with a variant of the neighbor-joining method, BIONJ (65). Because we did not use labor-intensive manual procedures to inspect multiple sequence alignment of each individual protein cluster and correct potential errors in automated annotations, the branch length might not be accurate. Our aim was to derive an unrooted tree with the correct topology in an automated fashion. Bootstrap was carried out with 100 random shuffling of the pairwise alignments. Only two branches had bootstrap values less than 100% (80% and 64% respectively). The derived tree was consistent with previous published results (63,66).
Calculation of reverse transcriptase footprints (RTF)
We define the RTF as the genomic fragments that after translation have significant sequence identity to known RT. The amount of RTF reflects RT activity. We first obtained 27,853 RT protein sequences from Swiss-Prot (April 2008 cutoff date). Then, we used the protein sequences to do blast search against all 16 fungal genomic sequences. The blast hits were noisy, and there might be multiple hits of RT in one region. We found that the two-dimensional chaining algorithm (67) was suited for sorting out the noisy RT hits in the genome. The algorithm was implemented in a program PFOG (Zhou unpublished) that first selected significant hits then build the “hit-chains” (footprints). Finally, footprints in the same area were consolidated: overlapping footprints will be combined. The number and length of each footprint were tabulated for each genome.
Relative intron location (RIL)
RIL is the nucleotide position of the intron in the mRNA divided by the total length of the mRNA. This is a value between 0 and 1 and usually stated as percentage; 0 means 5'-end and 1 means 3'-end.
Exon length distribution
Gene models from fungal genomes were mostly generated by protein homology and ab initio methods with varying degree of EST-models. For Chlamydomonas reinhardtii, we used 308,185 EST sequences and genomic sequences from DOEJGI (http://genome.jgi.doe.gov/Chlre4/Chlre4.home.html) as input for the COMBEST algorithm (K Zhou, manuscript in preparation) to derive EST-based gene models. Partial and single-exon gene models were excluded. Then unique coding exon length distribution was tabulated.
Simulation of intron loss
We first characterized intron number and exon length distribution for fungal genomes used in this study; both features can be closely represented by gamma distributions (with round up). Two parameters alpha (shape) and lambda (scale) determine the gamma distribution. To simulate ancestors with different average number of introns, we choose different (alpha, lambda) pairs, for example, (alpha =2.42, lambda =0.31) for introns per gene (IPG) 7.67, (alpha =2.297, lambda =0.255) for IPG 9.0. For exon length, we estimated the gamma parameters for each group of genes with 1, 2,…, 70 EPG based on the fungal genomes in this study. This forms a lookup table. The gamma parameters for exon length were then adjusted so that the protein length is around 400 aa. In this study, we picked the proper parameters to closely resemble the biological data. For the ancestor genome was represented by 10,000 genes that we simulated with random exon-shuffling. Essentially, exon length was generated from a gamma distribution with the number of exons controlled by another gamma distribution. No intron length was simulated.
For intron loss, we found that a linear probability distribution with density functions: f(x)=2(1-b)x+b, where 0≤b≤1 and x is RIL, gave the best results. The simulation was not very sensitive to b, and b in the range 0.5-0.8 worked well. A subset of genes seemed to be resistant to intron loss, which is represented by a pair of parameters (c, d): fraction of genes resistant to loss (c), and fraction of introns that are resistant to intron loss when the gene is resistant to intron loss (d). The parameter c ranges from 0.2-0.4, and d of 0.7-0.9 gave reasonable simulation results. The intron loss was simulated for 64 rounds with each round having the same number of RT-mediated intron loss events (simply referred as RT events). For looking at longer evolutionary distances, we use large values of RT events [4,000-8,000], and for looking at short evolutionary intervals we use smaller number of RT events [300-800]. Gene features: distribution of number of introns, exon length, and RIL were tabulated after each round.
Results
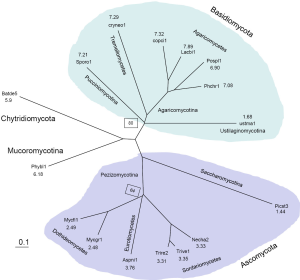
We characterized 16 fungal genomes (68) (Table 1) by constructing a phylogenetic tree and computing the average EPG from genes conserved in all genomes (Supplemental I; Figure S1). Representing Saccharomycotina, P. stipitis branched off from other Ascomycota clades. U. maydis, representative of Ustilaginomycotina, branched off from other Basidiomycota species. These two genomes experienced dramatic intron loss, but U. maydis lost significantly more than P. stipitis with respective to their immediate ancestor. EPG of Ascomycota was less than half that of Basidiomycota, and EPG of Chytridiomycota and Mucoromycotina was slightly lower than that of Basidiomycota. The amount of RT in each genome was measured by the amount of RTF and showed wide variations (Supplemental II; Figure S2). Although amount of RTF directly correlated with fugal genome size, the number of genes was a stronger determinant (Supplemental III; Figure S3).
RIL traces RT-mediated intron loss and links to ancestral intron number
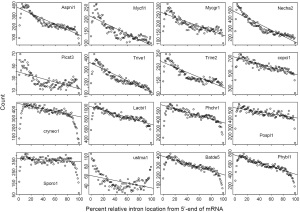
Difference of RIL in eukaryotic genomes has been noticed in a study of mRNA-mediated intron loss (52). Here we seek to link RT to intron loss through the variation of RIL (Figure 1) and difference of RTF (Supplemental II). There was a paucity of introns near both ends of mRNA (edge effects), which could be explained by random exon-shuffling as a mechanism for ancestral gene birth, as oppose to random insertion (Figure 2A at zero round). LECA could have achieved the intron-rich state, with the mean RIL being 0.5, through a variety of mechanisms (e.g., exon-shuffling or intron random insertion) either in a big bang fashion (69) or lack of RT-mediated intron loss in the initial period. As long as RT played a role in intron loss afterwards (70), the mean RIL would decrease with cumulative exposure to RT-mediated intron loss.
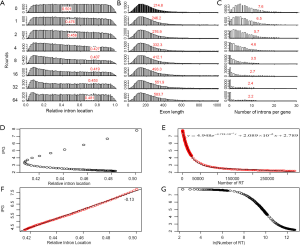
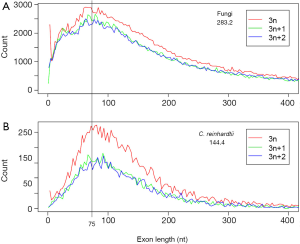
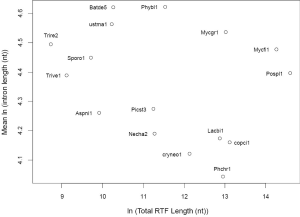
In order to understand the effect of RT on EPG, we first characterized the simulated intron loss (Figure 2 see Methods for details). The outcomes of the simulation were essentially the same with varying (IPG = EPG-1; from 6 to 15) in the ancestral genomes. In Figure 2, the ancestral genome had an average of 7.6 IPG. The most striking result was the preservation of peak length (mode) although the average exon length increased responsively with intron loss. This explained the similar modes of exon length distribution between C. reinhardtii and fungi (Figure 3) and laid a foundation to estimate the predominant exon length of LECA from modern genomes (Figure 4A; to be detailed in later section). Furthermore, the sparse introns near both ends of simulated genes concurred with both fungal (Figure 1) and other genomes (52).
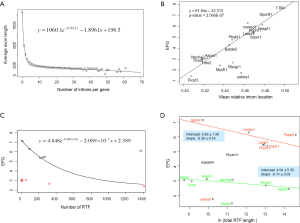
The relationship between IPG and RIL was an L-shaped curve with the top portion being almost linear in both simulation (Figure 2D,F) and real data (Figure 4B). The L-shaped curve is due to the conservation of introns at random locations in the gene (as introns may acquire function during evolution). The linear model, based on the top portion of the simulated data (excluding the ancestral data point), underestimated the ancestral IPG (–0.111±0.031, n=70) by a small margin. This is partly due to the heavy sampling at lower IPG. With even sampling, the underestimate shall be smaller. Excluding genomes near the bottom end of L-curve, M. graminicola and U. maydis, we arrived at 7.66 EPG for LECA although the data were exclusively fungi.
Missing RTF revealed by comparison between simulation and real data
The relationship between IPG and number of RT-mediated intron loss events (henceforth RT events) in simulation resembled exponential decay in an initial phase, followed by a linear phase, which can be described by a function y=4.948e–0.00004718x –0.000002089x+2.789 (Figure 2E). This equation stated that RT-mediated intron loss was less effective once intron distribution became 5’-biased. An RT event comprised two tandem random processes: (I) cDNA made from mRNA (catalyzed by RT) and (II) cDNA recombined with its cognate genomic DNA. RT events may or may not result in intron loss and could not be measured directly (RIL is a proxy). RTF can be measured and reflects the amount of RT, but the amount of RTF can be reduced by genomic DNA loss (Figure S2). On a whole genome level, total length and number of RTF were essentially interchangeable. Mammalian (71,72) and plant (73) genomes have abundant transposons, whereas most fungal genomes have very little (<10%) owing to genome-defense mechanisms and selection for smaller genomes (74-81). Therefore, RTF could be eliminated in fungal genomes at a faster rate. Moreover, one RT (represented by RTF) can catalyze more than one cDNA synthesis thus contributes to multiple RT events. Both of the above were reflected in the much faster decay rate for RTF (2.118×10–3 as compared to 4.718×10–5 for RT events) estimated from the Basidiomycota genomes by excluding unusual genomes (Figure 4C). Accordingly, Ascomycota had lost at least 1,400 RTF compared to Basidiomycota (data points need to be shifted to the right to fit the curve).
In simulation, more RT events led to more intron loss and less EPG. This was true within respective Ascomycota and Basidiomycota except for genomes that also lost the bulk of RTF. RT propagation is a growth process (exponential), and log transformation converts it to a linear variable. In simulation, the relationship between IPG and number of RT events in log scale was not linear (Figure 2G); however, within a phylum, the relationship between EPG and ln(tatol RTF length) was linear with negative slopes (Figure 4D). In consistent with simulation, Basidiomycota lost more introns per RTF (slope –0.30±0.16) than Ascomycota (slope –0.11±0.03). The result also agreed with previous observations: ten adjacent introns were lost in one successful RT event for Basidiomycota (50), but the highest was four in Ascomycota (48). Our result revealed the link between RT and intron loss (43,48,50,52,70,82) from an evolutionary perspective. U. maydis was an exception among the Basidiomycota genomes; its observable RTF [ln(tatol RTF length) =10.2] is far less than expected [ln(tatol RTF length)=26.9 from EPG of 1.75] according to the regression line for Basidiomycota (Figure 4D). The natural explanation was that the majority of RTF was removed after they had exerted their effect on intron loss. Less likely, RT-independent mechanisms accounted for the massive intron loss in U. maydis. There was still the possibility that RT in U. maydis was significantly more potent in catalyzing cDNA synthesis. The same could be stated for P. stipitis with respect to Ascomycota genomes.
The intercept of the regression line (Figure 4D), mathematically, denoted the EPG when total RTF Length =1 nt; biologically, it signified EPG in the presumptive ancestral genome when there was no RTF. Retroposons are dynamic (78,83) and go through cycles of boom-and-bust by interacting with the host defense system in fungi (84), but they are still useful for charting short term evolutionary histories of less than 150 Ma for mammals (85). If the ancestors for Basidiomycota and Ascomycota, respectively, had no active RT, then they had 9.69±1.99 EPG (P value 0.133), and 4.04±0.35 EPG (P value 0.026), respectively. The lack of RT in the ancestor of Basidiomycota could be deduced from the RIL (Figure 1); whereas, large scale RT loss from the ancestor of Ascomycota could be deduced from the comparison of simulation and real data. Nonetheless, if the respective ancestors for Basidiomycota and Ascomycota had one RTF (assuming one RT domain for about 1 kb), then they would have 7.6 and 3.3 EPG, respectively.
Deviation from the regression line contained information about the peculiarity of each genome’s response to RT and potentially other evolutionary events. Genomes below the regression line had accelerated intron loss owing to either more effective retroelements or selective pressure favoring intron loss. Expansion of genes coding for short peptides in L. bicolor (86) contributed to the average low EPG even though it had the highest EPG in highly conserved genes (Figures 5,S2). Genomes above the regression line were more resistant to intron loss, possibly, through one of the several genome-defense mechanisms such as repeat-induced point mutation (RIP) (81) or small RNA mechanisms (87). However, EPG of P. placenta might be slightly distorted upward owing to its diploid genome assembly and filtering procedures to derive the haploid gene set. The slight deviation in P. placenta and L. bicolor in EPG could contribute to the higher P value of Basidiomycota.
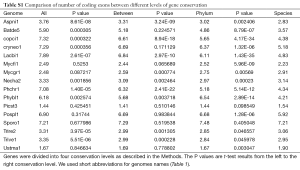
Full table
Genome-defense mechanisms may dampen the variation of RT-mediated intron loss. The data from Ascomycota had smaller variations compared to those from Basidiomycota. One explanation is the limited taxonomic distribution of genome-defense mechanisms in Basidiomycota (77,84). Diverse Ascomycota species had RIP (75,76,79,80,81,88).
Mucoromycotina and Chytridiomycota did not fit the regression lines of either Ascomycota or Basidiomycota (Figure 4D), which reflected the difference of each phylum’s ancestor. Finally, the effect of RT on intron length was not obvious (Supplemental IV; Figure S4). This is consistent with the proposed mechanism of RT-mediated intron loss where RT only interacts with mRNA, and cDNA only recombines with genomic DNA in exons.
Ancestral EPG estimated from dominant ancient exon and protein lengths
Since the mode of exon length distribution is conserved despite intron loss (Figure 2B), extant genomes still contain the information about the predominant exon length in the eukaryotic ancestor. Not surprisingly, the exon length distributions of fungi and green algae C. reinhardtii shared a peak around 75 nt (Figure 3). The latter genome exhibited features of the ancestor of both plant and animal (34). Both fungi and green algae are lower eukaryotes and are considered to be more close to ancestral life forms than higher eukaryotes. Ancient exons are usually flanked by short introns in mammalian genomes (89), but this criterion would not help us find ancient exons in fungi because most fungal introns are short. Another ancientness indicator, evolutionary conservation, is also not helpful because it implies ancientness only at protein level not exon/intron structure (90). If intron loss dominates eukaryotic evolution, then genes with the least intron loss must contain the most ancient exons. This means that genes with the greatest number of introns contain the most ancient exons. However, the number of genes with the greatest number of introns was small and had large variations. Therefore, we investigated the relationship between the average coding exon length (ACEL) and IPG from all 16 genomes and found an equation relating the two (Figure 4A). The right-hand side of Figure 4A represented the ancestral state of exon length of 66-86 nt. Seven proteins with 50 or more introns were all ancient (Supplemental V; Table S2). Coincidentally, the ancient protein module size is 25 aa (75 nt) (91) that is slightly longer than the 60 nt exon based on the theory of exons originating from random ORF (92). During novel gene-birth, stop codons mutated to sense codon, and frame-shifts disappeared by short deletions (93). Stop codon elimination mechanisms would be expected to be deployed in general exon birth process; therefore, the average exon length in the ancestor should be longer than 60 nt. With the above converging evidence, we concluded that the most prevalent length of ancient exons is 75 nt. However, this value is slightly smaller than the most frequent intron length of 90 nt from human, mouse, and C. elegans; nonetheless, all three genomes have prominent shoulders around 60 nt in the coding-exon length distribution curve (90).

Full table
Protein length is conserved in all eukaryotes (94), with median of 361 aa (95) (2004 survey) and 414 aa in a more recent survey of 215 genomes (96); the latter is close to the median 400 aa calculated from the 16 fungal genomes in this study. With 1,200 nt (400 aa) CDS length and 75 nt exon length, the EPG in the ancestor would be about 1,200/75=16.
Comparison of RIL in conserved and non-conserved genes
Comparing RIL between conserved and non-conserved genes can reveal whether they have been subjected to different extent of 3’-biased RT-mediated intron loss (Figure 6). All Ascomycota species had more 5’-introns in conserved genes compared to non-conserved genes with P value less than 2.0E-07 except for P. stipitis (with the least introns, P value 1.3E-03). This implies more RT-mediated intron loss during the early stage of Ascomycota evolution assuming similar processivities of RT’s in earlier and later time periods. The ancestor of Ascomycota might had lost ~4.4 EPG according to estimates in this paper, and only P. stipitis had further intron loss of lesser degree. Significant intron loss in the ancestor of Ascomycota was also reported by others (25,40). A brief period of dramatic loss of RT from Ascomycota ancestor might have followed the dramatic intron loss because of positive selection for a compact genome (97,98). This trend was similar but less dramatic in B. dendrobatidis (Chytridiomycota) and P. blakesleeanus (Mucoromycotina).
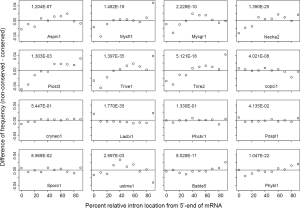
Four Basidiomycota genomes (C. neoformans, P. chrysosporium, P. placenta, and S. roseus) did not show differences between conserved and non-conserved genes, which indicated minimal and constant RT-exposure throughout their evolution history. In contrast, conserved genes of C. cinereus had more introns located near the 3'-end, which indicated more recent RT-mediated intron loss in younger genes (P value 4.02E-08). More introns of non-conserved genes in L. bicolor were located at both ends of genes (P value 1.8E-35); this was consistent with exon-shuffling at both ends of younger genes (99). The conserved genes of U. maydis lacked introns in the center and have excess introns in the 3'-end relative to non-conserved ones (P value 3.0E-03). For the U. maydis genome, conserved genes preferentially lost intron in the center, and younger genes preferentially lost introns in the 3' portion.
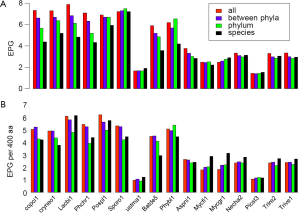
Fewer EPG in younger genes
The broad taxonomic distribution of a gene correlates with conserved protein sequences and gene structure, and is indicative of ancient origin. By comparing EPG at four different conservation levels: conserved in all species, between phyla, within a phylum, and species-specific (see Methods for detail), we found three types of genomes, with unchanged, decreasing, or increasing trend of EPG as conservation level went down. The majority of genomes showed a downtrend, whereas M. graminicola, with symptoms of hyper recombination (100), showed an upward trend (Figure 5A). At conservation levels “all”, “between phyla”, and “within phylum” U. maydis showed no difference (all had 1.7 EPG), but EPG at species level (1.9) was slightly larger and statistically significant (P value 0.003). S. roseus and P. stipitis showed no significant difference between conservation levels. Three genomes, N. haematococca, T. reesei, and T. virens, had very similar patterns where the first three conservation levels showed a downtrend and the species level showed a slight uptrend.
Although species-specific genes tended to have the smallest EPG, their exon densities were usually higher than those of genes conserved within phylum. Because protein lengths were shorter at lower conservation levels (data not shown), we normalized EPG to 400 aa, the median protein length in fungi (Figure 5B). The normalized value is hereby denoted as “exon density”. The general downtrend of exon density was similar to that of EPG for most Basidiomycota genomes, except for species-specific genes showing an uptrend, compared to phylum level, in four genomes: L. bicolor, P. chrysosporium, P. placenta, and S. roseus. The downtrend exon density had no parallel in EPG for S. roseus. The patterns of exon density and EPG were similar for B. dendrobatidis and P. blakesleeanus, as well as four Ascomycota genomes. The species-specific genes had more pronounced uptrend in exon density in five Ascomycota genomes: N. haematococca, M. graminicola, T. reesei, and T. virens. Although species-specific genes have fewer EPG they have higher exon density. This indicated much smaller exons in species-specific genes compared to genes conserved at phylum level.
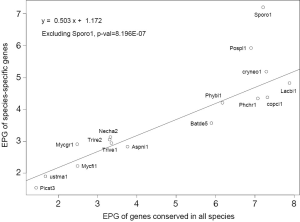
The general trend of fewer numbers of exons in less conserved genes could be quantified by a linear equation relating EPG of “all species” (x) and species-specific (y; Figure 7): y =0.503x +1.172, with P value 8.2E-07. The only exception was S. roseus, where EPG did not change in different conservation levels. This speaks to both intron loss and the difference between the ancient and modern gene birth processes.
Discussion
The knowledge of intron-rich eukaryotic ancestor has significantly changed our understanding of evolution of eukaryotic gene structure. More accurate assessment of EPG in the LECA anchors reasoning about early gene structure evolution. Previous methods of studying intron evolution relied on well-curated and broadly conserved orthologous genes and treated introns as binary character states in the context of protein sequence alignments (25,28,31,37,39,40). Such methods are powerful (60) but more sophisticated and have been thoroughly evaluated (101). Guided by a simple intron loss model, we have examined intron number evolution by comparing simulated results to observations from 16 fungal genomes and by seeking intra-genomic trends. The ancestral genome for the simulation contains genes that are assembled by random exon shuffling (Figure 2A). The distribution of RIL of this ancestor resembles that of S. roseus with intron avoidance at both ends (Figure 1). This model also assumes random conservation of a small fraction of the introns and a 3'-bias for RT-mediated intron loss. Although RT is very dynamic in genomes, RIL serves as a faithful marker for RT-mediated intron loss (Figures 2D,F,4B). The simulated relationship between EPG and RIL assumes an L-shaped curve that is observed in fungal genomes. Using this relationship, we arrived at 7.7 EPG in the LECA even though this value is biased because the data are exclusively fungal genomes. The presumptive ancestor with RIL being 0.5 implicates LECA. This view is corroborated by earlier results where the last fungal common ancestor (LFCA) lost introns after descending from Opisthokont (25,40) that is a descendant of LECA and gained 0.8 intron (40). Intron loss from LFCA is more likely mediated by RT and reduces the RIL. The RIL method underestimates ancestral EPG given the existence of none 3'-biased intron loss mechanisms: higher recombination rate in the middle of the gene (43,48), random deletion (generally accepted but no published evidence), or others (102).
Beyond the RIL method, exon length provides another channel for seeking the intron density of the ancestral genome. Underlying the estimate of 16 EPG in LECA is the conservation of most frequent exon length that is observed both in simulation and divergent genomes (green algae and fungi). Several lines of evidence converge on a single value of about 75 nt: (I) ancient protein module size of 25 aa (20,92,103,104); (II) exons originating from random ORF (92); (III) ancient exon length of fungi (Figure 4A); (IV) shared most frequent exon length from two primitive life forms (Figure 3). However, the mode of exon length is about 90 nt for three animal genomes (90). It is still a question whether animal genomes have selected for longer exons and longer proteins compared to fungi and green algae. The following observations favor this proposition: (I) evolution of long introns in animals, which correlates with younger exons (89) (II) splicing enhancers and suppressors in exons (11,105), in contrast to low information content in the exons of fungal genomes (24) (III) exon definition mechanism of splicing (51,106-109). In fact, this raises a very interesting possibility that fungal, particularly Basidiomycota, genomes harbor the most ancient exons, which is supported by the slightly shorter mode of exon length of fungi compared to that of C. reinhardtii (Figure 3) that inherited features from the common ancestor of both plants and animals (34).
Abundance of 3n length (multiples of 3) exons provides evidence for ancient exon-shuffling (Figure 3). Exon length of 3 nt is over represented in both fungal and C. reinhardtii genomes. This could be an ancestral feature. The over representation of 3n exons is more pronounced in C. reinhardtii compared to fungi, which could be the result of different annotation methods: predicted gene models dominate fungi, whereas the gene models from C. reinhardtii are exclusively EST-based. Symmetric exons (exons with the same intron phase on both sides) have been taken as evidence of exon-shuffling (99,110,111). Exons of 3n length could also benefit exon-shuffling during the ancient gene-birth process by maintaining reading frames. The 3n exons would be preferred if the shuffling process insert exon in between two exons (this is the case for modern exonization to be discussed later), or the shuffling involves random joining of exons. Furthermore, 3n exons could be the relics of ancient protein modules that could be translated into short peptides.
Why do two methods in this study predict very different ancestral EPG’s? The result from the RIL method is slightly larger than estimates from the classical methods: EPG (calculated from reported intron density and genes with average 1,200 nt ORF) 5.8 (25) and 6.7 (40) for LFCA that had loss introns after divergence from the common ancestor of Opisthokonts (25,28,40). However, the RIL method may give better estimates considering that classical methods could underestimate (50,112). Nonetheless, all the above methods (including the RIL method in this paper) suffer from heavy dependence on the intron states of contemporary genomes. The exon length method seeks the EPG of the very first intron-containing eukaryote that may well be the LUCA (113,114) and dominated by introns (80% genome) (115). Intron loss from the first intron-containing genome to that of LECA, if not mediated through RT, would be invisible to the RIL method. Furthermore, this early period may correspond to the hundreds of millions of years of early eukaryote evolution that is inaccessible to comparative approaches (116). Following this argument, 8.3 introns have been lost from the very early eukaryote to LECA (40,116), which is a very large number that still waits for further investigation.
The dependency of EPG on the amount of RTF is not a broadly useful method for estimating ancestral EPG, but it serves as a means to understand evolutionary events affecting EPG or RT. When used as a method to estimate ancestral EPG, none of the input genomes should have experienced significant change in selection for or against either intron loss or RT after divergence from a common ancestor. Furthermore, this method requires knowing the amount of RTF in the ancestor that is only known in special circumstances, such as the negligible amount of RTF in the ancestor of Basidiomycota. For individual genomes, deviation from the regression line between EPG and ln(tatol RTF length) signal either change in EPG or dynamics of RT (Figure 4D). For example, the expanding family of short genes in L. bicolor (86) contributes its being below the regression line. Transposons in fungal genomes can accumulate to large fractions (97,117-120) or can be lost to few active copies (74,121-123). P. placenta has the most RTF among Basidiomycota genomes; this may have caused its data point to shift to the right. U. maydis genome is shifted to the left by dramatic RTF loss. Finally, the dependence of EPG on RTF provides a long sought after evolutionary link between RT and intron loss (124) that has been proposed since 1985 (125,126), verified by cleaver molecular biology manipulations in yeast (127,128), and observed individually in fungal genomes (43,48,50).
In addition to serving a method of estimating ancestral EPG, the linear relationship between EPG and RIL divides genomes into those that are more sensitive or resistant to RT-mediated intron loss. For example, RT has a smaller effect on intron loss in C. neoformans that is above the regression line, which is consistent with earlier results (50). S. roseus is both above the regression line and closest to the ancestral state by all methods in this study and in intra genomic comparisons. This is consistent with the fact that S. roseus belongs to the Pucciniomycotina subphylum that is the only Basidiomycota group possessing the RIP genome-defense mechanism (77,84).
RT-mediated intron loss has both 3' and center bias in both Ascomycota (48) and Basidiomycota (43,50) genomes. In most genomes, the 3'-bias dominates, except for U. maydis where center bias is more prominent (Figure 1). Intra-genomic comparisons can reveal the relative timing of exposure to dramatic RT-mediated intron loss. The ancestor of Ascomycota lost significant introns (25,40). The excess of 5'-located introns in conserved relative to non-conserved genes Ascomycota genomes (Figure 6) suggests that RT plays a role in intron loss of the ancestor of Ascomycota. For most Basidiomycota genomes, there is no difference in RT exposure between early and later evolutionary times, but U. maydis has more intron loss from the center during early evolution (Figure 6). We interpret this as an early period of intron loss mediated by a highly processive RT (this period may coincide with the early divergence from the Basidiomycota ancestor); during this process, the ancestor of U. maydis lost most introns (25). The intron loss in younger genes is more 3'-biased. This also implies that the highly processive RT during early evolution was replaced later by a less processive RT. The current U. maydis genome had very little RTF (Figure S2).
Generally, more conserved genes (older) have more introns than less conserved ones (112,129) (Figure 5A). This can be attributed to either less conserved genes have lost more introns or the de novo gene-birth mechanisms produce genes with fewer introns; however, it is difficult to separate the two intertwined factors. Modern species-specific gene birth mechanisms tend to involve RT, which produces intronless or intron-reduced genes (129-131). Gene-birth through domain-shuffling (132) or trans-splicing (67) uses raw materials that have experienced intron loss although this process also creates one extra intron. If new genes were created by gene duplication followed by diversification, then they are more likely to lose introns to reduce the chance of homologous recombination among paralogs (133,134). De novo gene-birth through mutation of random sequences would definitely produce shorter genes with fewer introns (93). Several recent reports show that the strictly species-specific de novo genes usually have one and at most four exons (135-138), even in intron rich species such as human (139,140), mouse, and rat (141). Species-specific genes in Aspergillus cluster into regions enriched for transposons, have fewer introns, and are shorter (142); this agrees with our results (Figure 5A). However, the trend for exon density differs from that for EPG; particularly, species-specific genes not only have fewer exons but also have higher exon density for most genomes (Figure 5B). This apparent paradox can be explained by species-specific genes having both shorter CDS and shorter exon (data not shown), which is a hallmark of de novo gene-birth. As an example, the higher exon density in species-specific genes relative to genes conserved at phylum level in L. bicolor is attributed to de novo exon creation in species-specific gene family expansion in this genome (86). The lack of de novo gene-birth can explain why a few genomes, C. cinereus, C. neoformans, B. dendrobatidis, P. blakesleeanus, and P. stipitis, do not show higher exon density in species-specific genes (Figure 5).
The shorter de novo exons in fungi have parallels in metazoan genomes where novel exons are born of exonization (143,144) (there is an explosion of reports in this area, and we are not going to list all of them). Furthermore, novel exons arise from de novo gene birth (135-139,141,145). Novel exon birth through exonization tends to go through alternative splicing (143,146,147) and generally produces shorter exons (most reports ignore the exon length statistics and usually list a few examples) (148-150). The shorter exon length of alternative exons has been known since 1995 (151) and validated later with large sample size (152). Although exonized Alus are about 10 nt longer than their non-exonized ancestor (150), they are still about 12 nt shorter than major isoform (older) exons (89). However, the genome environment in human seems to favor novel exons longer than 80 nt and of 3n length (153). The mean coding exon length is 149, 149, 152, 364, 365, and 210 nt in human, mouse, Ciona intestinalis, Drosophila melanogaster, Anopheles gambiae, and Caenorhabditis elegans, respectively (90). The relatively shorter novel exons from exonization are 87, 70, and 22 nt respectively for human, mouse, and zebrafish, respectively (154). The exonization process resembles ancient gene birth in (I) using more or less random sequences; (II) have to join exons together; (III) favor 3n length. They differ in several aspects: (I) TE dominates modern exonization process (155) because of their abundance and harboring consensus splice sites (156); (II) several mutational steps are needed for modern exonization whereas, the ancient gene birth process may have used shorter exons and occurred in a very short time; (III) alternative splicing may not have existed at the time of ancient gene birth but is linked to modern exonization (155,157); (IV) large portion of modern exonization may have regulatory role (158) as opposed to protein-coding in ancient gene-birth. The novel exons from the modern de novo gene-creation process are dominated by single exon genes (138-140) that have gone through several mutations to give birth to longer coding exons. For three human de novo genes, the mutations have lengthened the ancestral ORF by 2-3 folds (139) with resulting exons (363-489 nt), longer than the average exons of vertebrate (137 nt). In another study, 27 human de novo genes (all single exon) have an average length of 521 nt that are twice the cognate ancestral non-coding ORF length (140). It seems that the path of evolution of these de novo genes, collected by a criterion excluding repetitive sequences (including TEs) (136,139-141), is unlikely to lead to the intron-rich gene repertoire of the ancestral genome. In a rare case involving Alu in human, a de novo gene has six exons with only exon three and four containing the CDS and three exons of length 122, 146, and 149 nt (93) that are roughly twice the length of ancient exon length of 75 nt but significantly shorter than the average from the whole genome (159). However, higher proportion of de novo genes from protozoans has introns (136) (although few and short with average protein length of 106 aa), and CDS exons of de novo genes of both multi-exon (mean 159 nt, median 124 nt, and n=25) and single-exon (mean 381 nt, median 321 nt, and n=15) are significantly shorter than the whole genome average of 957 nt in the primitive eukaryote Plasmodium vivax with average 2.5 EPG and 192 nt introns (160). The peak exon length from novel multi-exon genes from P. vivax is about 60 nt (very small sample size) that is slightly shorter than the peak ancient exon length of 75 nt. The median exon length or multi-exon novel genes from P. vivax is very close to 121 nt, the median exon length of internal alternative exons in human (157). Therefore, it is plausible that both de novo and ancient gene birth process create shorter exons and long introns (Supplemental VI; Figures S5,S6) (89), which is consistent with longer introns having higher potential for exonization (156). However, the novel exons from both exonization and de novo gene birth in metazoan seem to be longer than the ancient exon length, which may be due to different constraints, available raw materials, and genomic environments. Awaiting is a more rigorous statistical analysis of exon lengths from exonization, de novo gene birth, and ancient exons from diverse phyla.
Although RT is linked to intron loss, it is not related to intron length (Supplemental IV). Furthermore, EPG has no apparent relationship with intron length; however, intron loss is linked to intron length variation at different conservation levels within genomes (Supplemental VI). Most fungal genomes contain old and novel introns that differ in their time of birth, rate of intron length reduction, and initial length. Novel introns have longer initial length, but have faster rate of shortening. After massive intron loss, the fungal genomes will be dominated by element-bearing introns that are longer than old introns but will not change their length over time. U. maydis and P. stipitis genomes are good examples where novel introns are much shorter than element-bearing introns.
One important distinction between eukaryotes and prokaryotes is the presence and absence of introns in the genome even though Archaea has eukaryotic-like information processing, protein synthesis, and secretion system. The enormous diversity of eukaryotic life forms is evolved from the single cellular LECA with multiple cases of evolution via secondary simplification of complex ancestor. Naturally, one would think that the high intron density in the earliest eukaryotic genome endowed the possibility of current diversity. Similarly, the contribution of introns to the process of a stem cell developing into a complex organism would not be far-fetched. Here we list a few examples where introns play a role in stem cell regulation. The transcriptional regulator NANOG and (161) and Protein Kinase C delta (162) have alternative splicing. The first intron of PRMT1 harbors the activator binding site for c-Myc upon activation by T3 during intestinal stem cell development (163). The Smad2 master regulator for the Nodal/Activin controlled embryonic stem cell fate decision itself has multiple alternative spliced forms, and the majority of DNA regulatory elements in the target genes are located to introns (164). Non-coding RNAs have been established as important regulators for embryonic stem cell maintenance and pluripotency (165), about one third of them have been located in introns during embryonic stem cell neural differentiation (166). A splicing factor, TEG-1 CB2BP2 regulates stem cell proliferation and sex determination in the C. elegans (167).
Conclusions
RT plays a role in intron loss of fungal genomes. Intron loss from the ancestor of Ascomycota is RT-mediated with more 3'-bias than subsequent intron loss. Although the amount of RTF in fungal genomes is dynamic (significant number of RTF has been lost from the ancestor of Ascomycota), RIL is a reliable tracer for RT-mediated intron loss. Accordingly, the LECA has 7.7 EPG that is slightly larger than 6.4 EPG from previous estimates. The mode of exon length distribution is conserved during RT-mediate intron loss, which leads to ~16 EPG for the very first eukaryotic genome. Both intron loss and novel gene-birth process contribute to fewer EPG in less conserved genes. Novel gene birth process tends to produce shorter exons and longer introns which results in higher exon density in species-specific genes compared to genes conserved in all species. Novel introns are longer than old ones, and element-bearing introns are the longest. Massive intron loss causes the enrichment of element-bearing introns. RT contributes to genomes size, but genomes size is not related to EPG.
Additional data and files
Supplemental materials:
- Phylogenetic tree and average EPG of genes conserved in all genomes;
- Characterization of RT footprints (RTF) in fungal genomes;
- Amount of RT as a co-determinant of genome size;
- Relationship between intron length and amount of RTF;
- Genes with large number of exons tends to be more ancient;
- Intron length variation at different conservation levels determined by the proportions of three types of introns: old, novel, and element-bearing.
Phylogenetic tree and average coding exons per gene (EPG) of genes conserved in all genomes
To assist understanding of intron number evolution, we constructed a phylogenetic tree from 288 clusters containing exactly one protein from each genome (Figure S1). The average for EPG was computed from 491 clusters that contained at least one gene from each genome. The tree agreed with known fungal taxonomy with Chytridiomycota and Mucoromycotina (both with uncertain affinities) more related to each other than to Ascomycota and Basidiomycota.
Within Basidiomycota, U. maydis branched off early from the rest (EPG from 6.9 to 7.9), and its low EPG (1.7) indicated loss of at least five introns. Branched off immediately afterwards, S. roseus conserved the most introns among all genomes.
In contrast, Ascomycota phylum had much lower EPG, which hinted a loss of around three introns from the ancestor of this clade. P. stipitis branched off early from the rest and might have lost at least one intron, which resulted in its lowest EPG (1.4). The Dothideomycetes class had about 2.5 EPG that is slightly lower than Sordariomycetes (3.8 EPG) and Eurotiomycetes (3.3 EPG).
B. dendrobatidis representing the Chytridiomycota phylum and P. blakesleeanus representing the Mucoromycotina subphylum had very similar EPG (around 6). Both genomes also had similar relative intron location (RIL) and other features (data not shown).
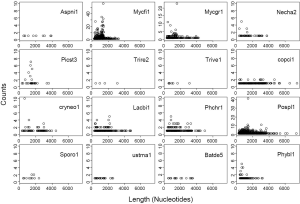
Characterization of RT footprints (RTF) in fungal genomes
An RTF is defined as the genomic sequences with significant similarity to proteins that contain RT protein domains (Methods for detail). These proteins usually contain other domains such as RNase-H, integrase, and maturase (168). Each RTF in a genome was characterized by two variables: count (copy number) and length (Figure S2). Long RTF indicates the presence of potentially active RT. High count indicates high RT activity either in the recent past or present. The two variables are different but not entirely independent. However, to characterize a genome with RTF we use either total count or total length of all RTF in the genome. The sum of the two variables are related by the average length of RTF (total length = total count × average length). This may explain our observation that total count and total length were interchangeable when studying relationships with other variables. For brevity, we presented results using either variable that was more meaningful in the context and used RTF to refer to the total amount in the genome without further clarification.
RTF was highly variable among fungal genomes. M. fijiensis and P. placenta had the most RTF count which is consistent with their genome expansion. C. cinereus and P. placenta contained the longest RTF’s. Closely related to M. fijiensis and also a plant pathogen, M. graminicola had moderate numbers of RTF, which was consistent with whole-genome analysis where eight dispensable chromosomes, moderate repetitive sequences, and RT’s had been subjected to repeat-induced point mutation (RIP) (100). The higher RTF in M. fijiensis compared to M. graminicola parallels the significant difference of repeat contents from a 100 kb region between the two species (169). The result for A. niger agrees with previous RT evaluation (170). T. reesei had the lowest RTF as well as the least transposable elements (171). Four Basidiomycota genomes: C. cinereus, C. neoformans, L. bicolor, and P. chrysosporium had moderate amounts of RTF. Two basal Basidiomycota genomes, S. roseus and U. maydis, had very few RTF’s. The former received no independent genomic analysis, but the later had. U. maydis is devoid of repetitive DNA and has a mechanism to prematurely terminate the transcription of foreign DNA (172). Worth noticing, two genomes P. stipitis and U. maydis with the most intron loss did not have the most number of RTF. Therefore, there is no straightforward correlation between the RTF and intron loss. Finally, the agreement between RTF and detailed characterizations of TE in the literature further validates the reliability of our procedure.
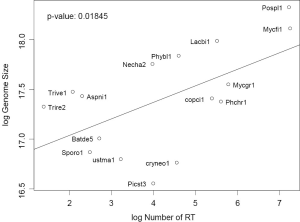
Amount of RT as a co-determinant of genome size
Genomes are mainly populated by protein-coding genes, transposons, and their relics (pseudogenes and disabled transposons), whereas rRNA and tRNA genes represent only a minute constant fraction. Not surprisingly, genome size can be explained by a linear model of the number of protein-coding genes and number of RTF: z =3322.2x +36718.4y –1513287.8 with P value 1.511e-07 where x, y, and z corresponds to number of genes, number of RTF, and genome size respectively. The parameter for x means the average genomic size per gene. The parameter for y means the average contribution to genome size (more than 7× the size of ORF for one typical fungal RT around 4.5-5 kb) by one RTF. Having multiple domains, RT proteins are longer than the average size of cellular proteins. Moreover, RT tends to pile up on older RT elements, giving rise to longer footprints. Our result hinted the fragmented nature of RTF in fungal genomes.
Gene number is more consistent than RT as a predictor of genome size. Most genomes (even with two extra fungal genomes not used in this study) were very close to the regression line linking genome size and gene number, except for genomes with the highest RTF: P. placenta and M. fijiensis (Data not shown). But most genomes were scattered away from the regression line between genome size and RTF (both variable in log scale) with P value =0.018 (Figure S3). This is due to the variation of both gene number, other types of transposable elements, and repetitive DNA sequences among fungal genomes.
Relationship between intron length and amount of RTF
In large genomes, transposable elements are mostly located in non-protein-coding regions (173). Long retroelements avoid genic regions, whereas short retroelements are enriched inside genes (174). There is a strong selection against LINE near protein-coding genes of human and mouse genomes (175) as well as in fungi (176,177). Accordingly, amount of RTF should not correlate with intron length in fungal genomes because the mostly short introns leave no room for retroelements. Indeed, we did not see any correlation between the two variables (Figure S4). However, a negative correlation became clear if we divided the genomes into two groups: the bottom group consisted of T. virens, T. reesei, S. roseus, A. niger, P. stipitis, N. haematococca, C. neoformans, L. bicolor, C. cinereus, and P. chrysosporium; the upper group comprised B. dendrobatidis, U. maydis, P. blakesleeanus, M. graminicola, M. fijiensis, and P. placenta. The two correlations had the same slope but different intercepts. One explanation is that genome expansion caused by RT activity triggers mechanisms for genome size reduction (genome defense) (178), including shortening of introns and loss of RTF themselves (cause for different intercepts). Being one of the genome defense mechanisms, repeated induced point mutation (RIP) that operates to different extents in fungal genomes (81) does not reduce genome size on the surface, but its G+C to A+T-rich effect could trigger deletions. Since most fungal genomes are limited in size, genomic DNA deletion must be a balancing force against transposons, which has been demonstrated in plant (179,180).
Genes with large number of exons tends to be more ancient
Ancient genes are intron rich (31), furthermore, they resist intron loss because all realistic intron loss mechanisms require homologous recombination that disfavor short exons. After examined seven genes with the most number of exons (50 or more), all from Basidiomycota, we found that all encoded ancient functions (Table S2). These genes preserved the ancestral short exon length by having little intron loss, but the number of exons might not represent ancient genes because of possible exon shuffling or gene fusion. Pospl1_99516 (the number is the JGI protein id; the prefix is the database name) might be a gene model chimera. Lacbi1_300923 (57 exons) and Pospl1_128265 (59 exons) were orthologs from L. bicolor and P. placenta, sharing 50 introns and 50% (70% non-gapped) protein sequence identity. Six intron of Lacbi1_300923 and eight introns of Pospl1_128265 were unique to each gene respectively, which lead to 74 introns in the ancestral gene (assuming intron loss dominates). Lacbi1_300923 had two EST supports at disjoint regions covering nine exons, which also indicated alternative splicing; the remaining exons were supported by sequence similarity to a predicted gene from C. cinereus that was an ortholog but had experienced intron loss. Pospl1_128265 was supported by EST at the 3'-end and had signs of alternative splicing by means of protein sequence identity. This protein contained an ancient histidine kinase domain found in eubacteria, archaebacteria, and eukaryotes (181) and could play a regulatory role in fungal development (182). Regulatory genes are usually expressed at very low levels and thus less likely to lose intron through RT-mediated pathway shown to be the primary means of intron loss (43,48,50). Two paralogs from P. placenta Pospl1_97544 and Pospl1_104193 shared 36 introns with 14 introns unique to each gene respectively and 74% (90% non-gapped) protein sequence identity. Compared with the ortholog pair, the paralogs pair shared more amino acid sequence identity but fewer common introns. The paralogs pair also lost the same number of introns. There appeared to be more intron loss in paralogs; this could be beneficial in reducing the chance of homologous recombination between paralogous genes. Pospl1_97544 had extensive EST support at various locations of the gene, but Pospl1_104193 had no EST support (one EST fell in the 3' half of the gene with different exon structure). Both paralogs showed extensive sequence conservation of exons with an orthologous genomic region of P. chrysosporium. This protein contained an alpha-isopropylmalate synthase domain (183), which was ancient and essential for very early life forms.
Sporo1_28096 had 58 exons and was a hypothetical protein. Phybl1_61381 had 62 exons and contained both cytochrome-C heme-binding site and PIKinase domain which is ancient being necessary for the emergence of cellular life. Both models had no protein or EST support. Phybl1_61381 had strong exonic genomic DNA sequence conservation with an orthologous region from R. oryzae.
Intron length variation at different conservation levels determined by the proportions of three types of introns: old, novel, and element-bearing
Intron length is determined by two opposing processes: elongation and shortening, and both are dynamic. Unequal cross-over during meiosis can both elongate and shorten introns, so it is the selection process that determines whether longer or shorter allele will dominate the population (184). If an intron contains sequence elements that increase the fitness of the organism, then the longer version would survive (185). However, even in the absence of apparent benefit, introns can get longer (186). DNA loss that has been deduced from transposon dynamics (178) can shorten the introns, whereas their expansion can occur by transposon insertions (174,187) that have not been observed in fungi (176,177,188). Majority of the fungal introns were short with 76% of them ranging between 50 and 200 nt (data not shown), in contrast to introns in plant and animal genomes. Furthermore, transposons in fungi generally avoid protein-coding genes (176,177) and only limited cases of enrichment toward 5'-region of subset of genes (122,188). Therefore, there had been a selection for shorter introns in fungal genomes.
The above conclusion might be the precursor for the general trend that more conserved genes have shorter introns than species-specific ones (Figure S5). However, this rule was not observed in all genomes, which became clear if we only look at the difference between the two extreme conservation levels (Figure S6A). The differences were statistically significant, except for A. niger, although the differences were small for several genomes, such as N. haematococca (Necha2) 3.7 nt and P. chrysosporium (Phchr1) 2.5 nt. Two genomes, P. stipitis and U. maydis, showed the opposite trend.
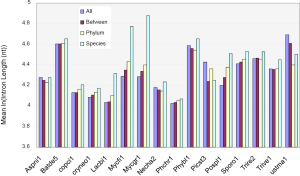
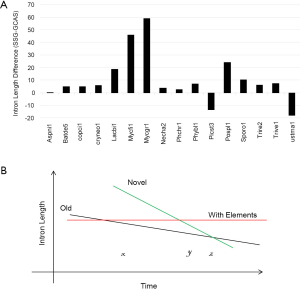
We could not find any apparent relationship between EPG and intron length. How are intron loss related to the variation of intron length within genomes? The answer is the random nature of the de novo gene birth process (143,189,190) in which transposable elements contribute to a larger fraction because of their abundance in contemporary genomes (156). Exons born from random sequences have an average about 60 nt (92); but introns would be much longer according to a simple model in which the average length of an intron arriving from random sequence is the inverse of the probability of sequence GT AG and associated consensus sequences such as branch points and pyrimidine tracks in proper locations (24). The distance between GT and AG must be greater than certain value ~40 nt for fungi, they both have to occur in the proper coding frame (for introns in coding regions), and be bordered by two exons. The minimal estimates is 256×3=768 nt that is much longer than the most frequent intron length of 55 nt in fungi (data not shown). Indeed, younger genes have longer introns in mammalian genomes (89), and species-specific genes have longer introns (Figure S5).
Under the selection for smaller genomes, old introns have longer time but slower rate to shorten themselves compared to novel ones (Figure S6B). Novel introns also have longer starting length than old introns. After massive intron loss, the remaining introns have higher chances of containing regulatory elements, are longer, and will not change their length afterwards. Each genomes have a particular combination of the above three populations of introns; however, technically it is difficult to separate them. In this study, species-specific genes had the highest proportion of novel introns, and genes conserved in all species had the least. Novel intron may be continuously introduced into the genome, or there may be bursts. Novel introns may differ in ages because “novel” means not shared by any genomes from a different species. Old introns have experienced evolution toward the optimal length. Corresponding to time x in Figure S6B were M. fijiensis and M. graminicola with old and novel introns; their species-specific genes had the longest introns (Figure S5) and highest exon density (Figure 5B), and their RTF were also the highest among Ascomycota (Figure S2). M. fijiensis also had the largest genomes among Ascomycota. These genomes had just experienced a genome expansion and de novo gene birth; genome reduction pressure has not yet acted upon the novel introns. The A. niger genome corresponded to time point z in Figure 5B where novel introns and old introns had similar lengths. There is no indication of significant de novo gene birth in the recent past in A. niger as marked by higher exon density in species-specific genes (Figure 5B) and very little RTF (Figure S2). U. maydis had the longest introns in genes conserved in all species. The introns in genes conserved in all species for P. stipitis are also the longest among Ascomycota genomes. U. maydis and P. stipitis genomes correspond to time point y in Figure S6B with novel and element-bearing introns. Both genomes have experienced genome size reduction and massive intron loss. U. maydis lost a lot more introns than P. stipitis relative to their respective ancestors (Figure S1); therefore, the former element-bearing introns population had a longer average length that the latter (Figure S6A). The longest average introns is found in S. cerevisiae genome where most introns are lost (228 introns left) (191). Most other fungal genomes would fall between time point x and z with mainly old and novel introns.
Acknowledgements
We thank Mingkun Li for his assistance in statistical and mathematical analysis, and Zhong Wang for critical reading and suggestions of the manuscript. Lixin Zhou from Pathway Genomics helps with a major revision.
Funding: The work conducted by the U.S. Department of Energy Joint Genome Institute is supported by the Office of Science of the U.S. Department of Energy under Contract No. DE-AC02-05CH11231.
Conflicts of Interest: All work, including major revisions of the manual script, for this paper by author K Zhou was done prior to joining BMS. The current affiliation with BMS is used for contact purpose only. The other authors have no conflicts of interest to declare.
References
- Rose AB, Elfersi T, Parra G, et al. Promoter-Proximal Introns in Arabidopsis thaliana Are Enriched in Dispersed Signals that Elevate Gene Expression. Plant Cell 2008;20:543-51.2.
- Bianchi M, Crinelli R, Giacomini E, et al. A potent enhancer element in the 5'-UTR intron is crucial for transcriptional regulation of the human ubiquitin C gene. Gene 2009;448:88-101. [PubMed]
- Fong YW, Zhou Q. Stimulatory effect of splicing factors on transcriptional elongation. Nature 2001;414:929-33. [PubMed]
- Reed R, Hurt E. A Conserved mRNA Export Machinery Coupled to pre-mRNA Splicing. Cell 2002;108:523-31. [PubMed]
- Kerényi Z, Merai Z, Hiripi L, et al. Inter-kingdom conservation of mechanism of nonsense-mediated mRNA decay. EMBO J 2008;27:1585-95. [PubMed]
- Mekouar M, Blanc-Lenfle I, Ozanne C, et al. Detection and analysis of alternative splicing in Yarrowia lipolytica reveal structural constraints facilitating nonsense-mediated decay of intron-retaining transcripts. Genome Biol 2010;11:R65. [PubMed]
- Mendell JT, Sharifi NA, Meyers JL, et al. Nonsense surveillance regulates expression of diverse classes of mammalian transcripts and mutes genomic noise. Nat Genet 2004;36:1073-8. [PubMed]
- Niu DK. Protecting exons from deleterious R-loops: a potential advantage of having introns. Biol Direct 2007;2:11. [PubMed]
- Irimia M, Rukov JL, Penny D, et al. Functional and evolutionary analysis of alternatively spliced genes is consistent with an early eukaryotic origin of alternative splicing. BMC Evol Biol 2007;7:188. [PubMed]
- Nilsen TW, Graveley BR. Expansion of the eukaryotic proteome by alternative splicing. Nature 2010;463:457-63. [PubMed]
- Chen M, Manley JL. Mechanisms of alternative splicing regulation: insights from molecular and genomics approaches. Nat Rev Mol Cell Biol 2009;10:741-54. [PubMed]
- Schwartz S, Meshorer E, Ast G. Chromatin organization marks exon-intron structure. Nat Struct Mol Biol 2009;16:990-5. [PubMed]
- Spies N, Nielsen CB, Padgett RA, et al. Biased Chromatin Signatures around Polyadenylation Sites and Exons. Mol Cell 2009;36:245-54. [PubMed]
- Kolasinska-Zwierz P, Down T, Latorre I, et al. Differential chromatin marking of introns and expressed exons by H3K36me3. Nat Genet 2009;41:376-81. [PubMed]
- Dhami P, Dhami P, Saffrey P, et al. Complex Exon-Intron Marking by Histone Modifications Is Not Determined Solely by Nucleosome Distribution. PLoS One 2010;5:e12339. [PubMed]
- Tilgner H, Nikolaou C, Althammer S, et al. Nucleosome positioning as a determinant of exon recognition. Nat Struct Mol Biol 2009;16:996-1001. [PubMed]
- Schwartz S, Ast G. Chromatin density and splicing destiny: on the cross-talk between chromatin structure and splicing. EMBO J 2010;29:1629-36. [PubMed]
- Cancherini DV, França GS, de Souza SJ. The role of exon shuffling in shaping protein-protein interaction networks. BMC Genomics 2010;11:S11. [PubMed]
- Elrouby N, Bureau TE. Bs1, a New Chimeric Gene Formed by Retrotransposon-Mediated Exon Shuffling in Maize. Plant Physiol 2010;153:1413-24. [PubMed]
- Liu M, Grigoriev A. Protein domains correlate strongly with exons in multiple eukaryotic genomes – evidence of exon shuffling? Trends Genet 2004;20:399-403. [PubMed]
- Vibranovski MD, Sakabe NJ. Souza SJd. A possible role of exon-shuffling in the evolution of signal peptides of human proteins. FEBS Lett 2006;580:1621-4. [PubMed]
- Gregory TR, Nicol JA, Tamm H, et al. Eukaryotic genome size databases. Nucleic Acids Res 2007;35:D332-8. [PubMed]
- Parenteau J, Durand M, Veronneau S, et al. Deletion of many yeast introns reveals a minority of genes that require splicing for function. Mol Biol Cell 2008;19:1932-41. [PubMed]
- Kupfer DM, Drabenstot SD, Buchanan KL, et al. Introns and splicing elements of five diverse fungi. Eukaryot Cell 2004;3:1088-100. [PubMed]
- Stajich JE, Dietrich FS, Roy SW. Comparative genomic analysis of fungal genomes reveals intron-rich ancestors. Genome Biol 2007;8:R223. [PubMed]
- Loftus BJ, Fung E, Roncaglia P, et al. The genome of the basidiomycetous yeast and human pathogen Cryptococcus neoformans. Science 2005;307:1321-4. [PubMed]
- Sverdlov AV. Conservation versus parallel gains in intron evolution. Nucleic Acids Res 2005;33:1741-8. [PubMed]
- Csurös M, Rogozin IB, Koonin EV. Extremely Intron-Rich Genes in the Alveolate Ancestors Inferred with a Flexible Maximum-Likelihood Approach. Mol Biol Evol 2008;25:903-11. [PubMed]
- Vanácová S, Yan W, Carlton JM, et al. Proc Natl Acad Sci U S A. 2005;102:4430-5. [PubMed]
- Slamovits CH, Keeling PJ. A high density of ancient spliceosomal introns in oxymonad excavates. BMC Evol Biol 2006;6:34. [PubMed]
- Roy SW, Gilbert W. Complex early genes. Proc Natl Acad Sci U S A 2005;102:1986-91. [PubMed]
- Nguyen HD, Yoshihama M, Kenmochi N. New Maximum Likelihood Estimators for Eukaryotic Intron Evolution. PloS Comput Biol 2005;1:e79. [PubMed]
- Rogozin IB, Sverdlov AV, Babenko VN, et al. Analysis of evolution of exon–intron structure of eukaryotic genes. Brief Bioinform 2005;6:118-34. [PubMed]
- Merchant SS, Prochnik SE, Vallon O, et al. The Chlamydomonas genome reveals the evolution of key animal and plant functions. Science 2007;318:245-50. [PubMed]
- King N, Westbrook MJ, Young SL, et al. The genome of the choanoflagellate Monosiga brevicollis and the origin of metazoans. Nature 2008;451:783-8. [PubMed]
- Roy SW, Penny D. Smoke Without Fire: Most Reported Cases of Intron Gain in Nematodes Instead Reflect Intron Losses. Mol Biol Evol 2006;23:2259-62. [PubMed]
- Roy SW, Gilbert W. Rates of intron loss and gain: implications for early eukaryotic evolution. Proc Natl Acad Sci U S A 2005;102:5773-8. [PubMed]
- Cho S, Jin SW, Cohen A, et al. A phylogeny of caenorhabditis reveals frequent loss of introns during nematode evolution. Genome Res 2004;14:1207-20. [PubMed]
- Carmel L, Wolf YI, Rogozin IB, et al. Three distinct modes of intron dynamics in the evolution of eukaryotes. Genome Res 2007;17:1034-44. [PubMed]
- Csuros M, Rogozin IB, Koonin EV. A Detailed History of Intron-rich Eukaryotic Ancestors Inferred from a Global Survey of 100 Complete Genomes. PLoS Comput Biol 2011;7:e1002150. [PubMed]
- Carmel L, Rogozin IB, Wolf YI, et al. A maximum likelihood method for reconstruction of the evolution of eukaryotic gene structure. Methods Mol Biol 2009;541:357-71. [PubMed]
- Brady SG, Danforth BN. Recent intron gain in elongation factor-1alpha of colletid bees (Hymenoptera: Colletidae). Mol Biol Evol 2004;21:691-6. [PubMed]
- Sharpton TJ, Neafsey DE, Galagan JE, et al. Mechanisms of intron gain and loss in Cryptococcus. Genome Biol 2008;9:R24. [PubMed]
- Roy SW, Fedorov A, Gilbert W. Large-scale comparison of intron positions in mammalian genes shows intron loss but no gain. Proc Natl Acad Sci U S A 2003;100:7158-62. [PubMed]
- Roy SW, Hartl DL. Very little intron loss/gain in Plasmodium: intron loss/gain mutation rates and intron number. Genome Res 2006;16:750-6. [PubMed]
- Roy SW, Penny D. Patterns of intron loss and gain in plants: intron loss-dominated evolution and genome-wide comparison of O. sativa and A. thaliana. Mol Biol Evol 2007;24:171-81. [PubMed]
- Fawcett JA, Rouzé P, Van de Peer Y. Higher intron loss rate in Arabidopsis thaliana than A. lyrata is consistent with stronger selection for a smaller genome. Mol Biol Evol 2012;29:849-59.
- Zhang LY, Yang YF, Niu DK. Evaluation of Models of the Mechanisms Underlying Intron Loss and Gain in Aspergillus Fungi. J Mol Evol 2010;71:364-73. [PubMed]
- Roy SW, Gilbert W. The evolution of spliceosomal introns: patterns, puzzles and progress. Nat Rev Genet 2006;7:211-21. [PubMed]
- Stajich JE, Dietrich FS. Evidence of mRNA-mediated intron loss in the human-pathogenic fungus Cryptococcus neoformans. Eukaryot Cell 2006;5:789-93. [PubMed]
- Niu DK. Exon definition as a potential negative force against intron losses in evolution. Biol Direct 2008;3:46. [PubMed]
- Niu DK, Hou WR, Li SW. mRNA-mediated intron losses: evidence from extraordinarily large exons. Mol Biol Evol 2005;22:1475-81. [PubMed]
- Bibiłło A, Eickbush TH. The reverse transcriptase of the R2 non-LTR retrotransposon: continuous synthesis of cDNA on non-continuous RNA templates. J Mol Biol 2002;316:459-73. [PubMed]
- Piskareva O, Schmatchenko V. DNA polymerization by the reverse transcriptase of the human L1 retrotransposon on its own template in vitro. FEBS Lett 2006;580:661-8. [PubMed]
- Galligan JT, Marchetti SE, Kennell JC. Reverse transcription of the pFOXC mitochondrial retroplasmids of Fusarium oxysporum is protein primed. Mob DNA 2011;2:1. [PubMed]
- Cristofari G, Gabus C, Ficheux D, et al. Characterization of Active Reverse Transcriptase and Nucleoprotein Complexes of the Yeast Retrotransposon Ty3 in Vitro. J Biol Chem 1999;274:36643-8. [PubMed]
- Hizi A. The Reverse Transcriptase of the Tf1 Retrotransposon Has a Specific Novel Activity for Generating the RNA Self-Primer That Is Functional in cDNA Synthesis. J Virol 2008;82:10906-10. [PubMed]
- Scofield DG, Hong X, Lynch M. Position of the final intron in full-length transcripts: determined by NMD? Mol Biol Evol 2007;24:896-9. [PubMed]
- Rogozin IB, Wolf YI, Sorokin AV, et al. Remarkable Interkingdom Conservation of Intron Positions and Massive, Lineage-Specific Intron Loss and Gain in Eukaryotic Evolution. Curr Biol 2003;13:1512-7. [PubMed]
- Carmel L, Wolf YI, Rogozin IB, et al. EREM: Parameter Estimation and Ancestral Reconstruction by Expectation-Maximization Algorithm for a Probabilistic Model of Genomic Binary Characters Evolution. Adv Bioinformatics 2010:167408.
- Csurös M. Malin: maximum likelihood analysis of intron evolution in eukaryotes. Bioinformatics 2008;24:1538-9. [PubMed]
- Smith TF, Waterman MS. Identification of common molecular subsequences. J Mol Biol 1981;147:195-7. [PubMed]
- Wang H, Xu Z, Gao L, et al. A fungal phylogeny based on 82 complete genomes using the composition vector method. BMC Evol Biol 2009;9:195. [PubMed]
- Grishin NV. Estimation of the number of amino acid substitutions per site when the substitution rate varies among sites. J Mol Evol 1995;41:675-9. [PubMed]
- Gascuel O. BIONJ: an improved version of the NJ algorithm based on a simple model of sequence data. Mol Biol Evol 1997;14:685-95. [PubMed]
- Fitzpatrick DA, Logue ME, Stajich JE, et al. A fungal phylogeny based on 42 complete genomes derived from supertree and combined gene analysis. BMC Evol Biol 2006;6:99. [PubMed]
- Gusfield D. Algorithms on Strings, Trees and Sequences: Computer Science and Computational Biology. New York: The Press Syndicate of the University of Cambridge; 1997.
- Cuomo CA, Birren BW. The fungal genome initiative and lessons learned from genome sequencing. Methods Enzymol 2010;470:833-55. [PubMed]
- Koonin EV. The Biological Big Bang model for the major transitions in evolution. Biol Direct 2007;2:21. [PubMed]
- Lin K, Zhang DY. The excess of 5' introns in eukaryotic genomes. Nucleic Acids Res 2005;33:6522-7. [PubMed]
- Lander ES, Linton LM, Birren B, et al. Initial sequencing and analysis of the human genome. Nature 2001;409:860-921. [PubMed]
- Smit AF. Interspersed repeats and other mementos of transposable elements in mammalian genomes. Curr Opin Genet Dev 1999;9:657-63. [PubMed]
- Schnable PS, Ware D, Fulton RS, et al. The B73 maize genome: complexity, diversity, and dynamics. Science 2009;326:1112-5. [PubMed]
- Nowrousian M, Stajich JE, Chu M, et al. De novo assembly of a 40 Mb eukaryotic genome from short sequence reads: Sordaria macrospora, a model organism for fungal morphogenesis. PLoS Genet 2010;6:e1000891. [PubMed]
- Hamann A, Feller F, Osiewacz HD. The degenerate DNA transposon Pat and repeat-induced point mutation (RIP) in Podospora anserina. Mol Gen Genet 2000;263:1061-9. [PubMed]
- Clutterbuck AJ. MATE transposable elements in Aspergillus nidulans: evidence of repeat-induced point mutation. Fungal Genet Biol 2004;41:308-16. [PubMed]
- Horns F, Petit E, Yockteng R, et al. Patterns of Repeat-Induced Point Mutation in Transposable Elements of Basidiomycete Fungi. Genome Biol Evol 2012;4:240-7. [PubMed]
- Neafsey DE, Barker BM, Sharpton TJ, et al. Population genomic sequencing of Coccidioides fungi reveals recent hybridization and transposon control. Genome Res 2010;20:938-46. [PubMed]
- Selker EU. Repeat-induced gene silencing in fungi. Adv Genet 2002;46:439-50. [PubMed]
- Ikeda K, Nakayashiki H, Kataoka T, et al. Repeat-induced point mutation (RIP) in Magnaporthe grisea: implications for its sexual cycle in the natural field context. Mol Microbiol 2002;45:1355-64. [PubMed]
- Galagan JE, Selker EU. RIP: the evolutionary cost of genome defense. Trends Genet 2004;20:417-23. [PubMed]
- Roy SW, Penny D. Widespread intron loss suggests retrotransposon activity in ancient apicomplexans. Mol Biol Evol 2007;24:1926-33. [PubMed]
- Gabriel A, Dapprich J, Kunkel M, et al. Global Mapping of Transposon Location. PLoS Genet 2006;2:e212. [PubMed]
- Johnson LJ, Giraud T, Anderson R, et al. The impact of genome defense on mobile elements in Microbotryum. Genetica 2010;138:313-9. [PubMed]
- Nishihara H, Maruyamab S, Okada N. Retroposon analysis and recent geological data suggest near-simultaneous divergence of the three superorders of mammals. Proc Natl Acad Sci U S A 2009;106:5235-40. [PubMed]
- Martin F, Aerts A, Ahren D, et al. The genome of Laccaria bicolor provides insights into mycorrhizal symbiosis. Nature 2008;452:88-92. [PubMed]
- Senti KA, Brennecke J. The piRNA pathway: a fly’s perspective on the guardian of the genome. Trends Genet 2010;26:499-509. [PubMed]
- Braumann I, van den Berg M, Kempken F. Repeat induced point mutation in two asexual fungi, Aspergillus niger and Penicillium chrysogenum. Curr Genet 2008;53:287-97. [PubMed]
- Roy M, Kim N, Xing Y, et al. The effect of intron length on exon creation ratios during the evolution of mammalian genomes. RNA 2008;14:2261-73. [PubMed]
- Yandell M, Mungall CJ, Smith C, et al. Large-scale trends in the evolution of gene structures within 11 animal genomes. PLoS Comput Biol 2006;2:e15. [PubMed]
- Fedorov A, Roy S, Cao X, et al. Phylogenetically older introns strongly correlate with module boundaries in ancient proteins. Genome Res 2003;13:1155-7. [PubMed]
- Regulapati R, Bhasi A, Singh CK, et al. Origination of the split structure of spliceosomal genes from random genetic sequences. PLoS One 2008;3:e3456. [PubMed]
- Li CY, Zhang Y, Wang Z, et al. A human-specific de novo protein-coding gene associated with human brain functions. PLoS Comput Biol 2010;6:e1000734. [PubMed]
- Wang D. A General Tendency for Conservation of Protein Length Across Eukaryotic Kingdoms. Mol Biol Evol 2005;22:142-7. [PubMed]
- Brocchieri L, Karlin S. Protein length in eukaryotic and prokaryotic proteomes. Nucleic Acids Res 2005;33:3390-400. [PubMed]
- Wang M, Kurland CG, Caetano-Anollés G. Reductive evolution of proteomes and protein structures. Proc Natl Acad Sci U S A 2011;108:11954-8. [PubMed]
- Spanu PD, Abbott JC, Amselem J, et al. Genome expansion and gene loss in powdery mildew fungi reveal tradeoffs in extreme parasitism. Science 2010;330:1543-6. [PubMed]
- Sahasrabuddhe VV, Bhosale RA, Kavatkar AN, et al. Comparison of visual inspection with acetic acid and cervical cytology to detect high-grade cervical neoplasia among HIV-infected women in India. Int J Cancer 2012;130:234-40. [PubMed]
- Vibranovski MD, Sakabe NJ, de Oliveira RS, et al. Signs of ancient and modern exon-shuffling are correlated to the distribution of ancient and modern domains along proteins. J Mol Evol 2005;61:341-50. [PubMed]
- Goodwin SB, M’Barek S B, Dhillon B, et al. Finished genome of the fungal wheat pathogen Mycosphaerella graminicola reveals dispensome structure, chromosome plasticity, and stealth pathogenesis. PLoS Genet 2011;7:e1002070. [PubMed]
- Cohen O, Pupko T. Inference of gain and loss events from phyletic patterns using stochastic mapping and maximum parsimony--a simulation study. Genome Biol Evol 2011;3:1265-75. [PubMed]
- Mitrovich QM, Tuch BB, De La Vega FM, et al. Evolution of yeast noncoding RNAs reveals an alternative mechanism for widespread intron loss. Science 2010;330:838-41. [PubMed]
- de Roos AD. Conserved intron positions in ancient protein modules. Biol Direct 2007;2:7. [PubMed]
- de Souza SJ, Long M, Schoenbach L, et al. Intron positions correlate with module boundaries in ancient proteins. Proc Natl Acad Sci U S A 1996;93:14632-6. [PubMed]
- Barash Y, Calarco JA, Gao W, et al. Deciphering the splicing code. Nature 2010;465:53-9. [PubMed]
- Sharma S, Kohlstaedt LA, Damianov A, et al. Polypyrimidine tract binding protein controls the transition from exon definition to an intron defined spliceosome. Nat Struct Mol Biol 2008;15:183-91. [PubMed]
- Ke S, Chasin LA. Context-dependent splicing regulation: Exon definition, co-occurring motif pairs and tissue specificity. RNA Biol 2011;8:384-8. [PubMed]
- Crabb TL, Lam BJ, Hertel KJ. Retention of spliceosomal components along ligated exons ensures efficient removal of multiple introns. RNA 2010;16:1786-96. [PubMed]
- Ke S, Chasin LA. Intronic motif pairs cooperate across exons to promote pre-mRNA splicing. Genome Biol 2010;11:R84. [PubMed]
- Patthy L. Intron-dependent evolution: preferred types of exons and introns. FEBS Lett 1987;214:1-7. [PubMed]
- Kawashima T, Kawashima S, Tanaka C, et al. Domain shuffling and the evolution of vertebrates. Genome Res 2009;19:1393-403. [PubMed]
- Carmel L, Rogozin IB, Wolf YI, et al. Evolutionarily conserved genes preferentially accumulate introns. Genome Res 2007;17:1045-50. [PubMed]
- Glansdorff N, Xu Y, Labedan B. The Last Universal Common Ancestor: emergence, constitution and genetic legacy of an elusive forerunner. Biol Direct 2008;3:29. [PubMed]
- Glansdorff N, Xu Y, Labedan B. The origin of life and the last universal common ancestor: do we need a change of perspective? Res Microbiol 2009;160:522-8. [PubMed]
- Koonin EV. Intron-dominated genomes of early ancestors of eukaryotes. J Hered 2009;100:618-23. [PubMed]
- Chernikova D, Motamedi S, Csuros M, et al. A late origin of the extant eukaryotic diversity: divergence time estimates using rare genomic changes. Biol Direct 2011;6:26. [PubMed]
- Kelkar YD, Ochman H. Causes and Consequences of Genome Expansion in Fungi. Genome Biol Evol 2012;4:13-23. [PubMed]
- Martin F, Kohler A, Murat C, et al. Perigord black truffle genome uncovers evolutionary origins and mechanisms of symbiosis. Nature 2010;464:1033-8. [PubMed]
- Ma LJ, Ibrahim AS, Skory C, et al. Genomic analysis of the basal lineage fungus Rhizopus oryzae reveals a whole-genome duplication. PLoS Genet 2009;5:e1000549. [PubMed]
- Parlange F, Oberhaensli S, Breen J, et al. A major invasion of transposable elements accounts for the large size of the Blumeria graminis f.sp. tritici genome. Funct Integr Genomics 2011;11:671-7. [PubMed]
- Espagne E, Lespinet O, Malagnac F, et al. The genome sequence of the model ascomycete fungus Podospora anserina. Genome Biol 2008;9:R77. [PubMed]
- Fleetwood DJ, Khan AK, Johnson RD, et al. Abundant degenerate miniature inverted-repeat transposable elements in genomes of epichloid fungal endophytes of grasses. Genome Biol Evol 2011;3:1253-64. [PubMed]
- Cuomo CA, Guldener U, Xu JR, et al. The Fusarium graminearum genome reveals a link between localized polymorphism and pathogen specialization. Science 2007;317:1400-2. [PubMed]
- Cohen NE, Shen R, Carmel L. The Role of Reverse Transcriptase in Intron Gain and Loss Mechanisms. Mol Biol Evol 2012;29:179-86. [PubMed]
- Fink GR. Pseudogenes in yeast? Cell 1987;49:5-6. [PubMed]
- Baltimore D. Retroviruses and retrotransposons: the role of reverse transcription in shaping the eukaryotic genome. Cell 1985;40:481-2. [PubMed]
- Derr LK, Strathern JN. A role for reverse transcripts in gene conversion. Nature 1993;361:170-3. [PubMed]
- Melamed C, Nevo Y, Kupiec M. Involvement of cDNA in homologous recombination between Ty elements in Saccharomyces cerevisiae. Mol Cell Biol 1992;12:1613-20. [PubMed]
- Fridmanis D, Fredriksson R, Kapa I, et al. Formation of new genes explains lower intron density in mammalian Rhodopsin G protein-coupled receptors. Mol Phylogenet Evol 2007;43:864-80. [PubMed]
- Xing J, Wang H, Belancio VP, et al. Emergence of primate genes by retrotransposon-mediated sequence transduction. Proc Natl Acad Sci U S A 2006;103:17608-13. [PubMed]
- Szcześniak MW, Ciomborowska J, Nowak W, et al. Primate and rodent specific intron gains and the origin of retrogenes with splice variants. Mol Biol Evol 2011;28:33-7. [PubMed]
- Kaessmann H, Zollner S, Nekrutenko A, et al. Signatures of domain shuffling in the human genome. Genome Res 2002;12:1642-50. [PubMed]
- Knowles DG, McLysaght A. High rate of recent intron gain and loss in simultaneously duplicated Arabidopsis genes. Mol Biol Evol 2006;23:1548-57. [PubMed]
- Katju V. In with the old, in with the new: the promiscuity of the duplication process engenders diverse pathways for novel gene creation. Int J Evol Biol 2012;2012:341932.
- Levine MT, Jones CD, Kern AD, et al. Novel genes derived from noncoding DNA in Drosophila melanogaster are frequently X-linked and exhibit testis-biased expression. Proc Natl Acad Sci U S A 2006;103:9935-9. [PubMed]
- Yang Z, Huang J. De novo origin of new genes with introns in Plasmodium vivax. FEBS Lett 2011;585:641-4. [PubMed]
- Cai J, Zhao R, Jiang H, et al. De novo origination of a new protein-coding gene in Saccharomyces cerevisiae. Genetics 2008;179:487-96. [PubMed]
- Carvunis AR, Rolland T, Wapinski I, et al. Proto-genes and de novo gene birth. Nature 2012;487:370-4. [PubMed]
- Knowles DG, McLysaght A. Recent de novo origin of human protein-coding genes. Genome Res 2009;19:1752-9. [PubMed]
- Wu DD, Irwin DM, Zhang YP. De Novo Origin of Human Protein-Coding Genes. PLoS Genet 2011;7:e1002379. [PubMed]
- Murphy DN, McLysaght A. De novo origin of protein-coding genes in murine rodents. PloS one 2012;7:e48650. [PubMed]
- Fedorova ND, Khaldi N, Joardar VS, et al. Genomic islands in the pathogenic filamentous fungus Aspergillus fumigatus. PLoS Genet 2008;4:e1000046. [PubMed]
- Sorek R. The birth of new exons: mechanisms and evolutionary consequences. RNA 2007;13:1603-8. [PubMed]
- Lin L, Jiang P, Shen S, et al. Large-scale analysis of exonized mammalian-wide interspersed repeats in primate genomes. Hum Mol Genet 2009;18:2204-14. [PubMed]
- Xiao W, Liu H, Li Y, et al. A rice gene of de novo origin negatively regulates pathogen-induced defense response. PloS one 2009;4:e4603. [PubMed]
- Sorek R. When new exons are born. Heredity (Edinb) 2009;103:279-80. [PubMed]
- Schmitz J, Brosius J. Exonization of transposed elements: A challenge and opportunity for evolution. Biochimie 2011;93:1928-34. [PubMed]
- Krull M, Brosius J, Schmitz J. Alu-SINE exonization: en route to protein-coding function. Mol Biol Evol 2005;22:1702-11. [PubMed]
- Lin L, Shen S, Tye A, et al. Diverse splicing patterns of exonized Alu elements in human tissues. PLoS Genet 2008;4:e1000225. [PubMed]
- Schwartz S, Gal-Mark N, Kfir N, et al. Alu exonization events reveal features required for precise recognition of exons by the splicing machinery. PloS Comput Biol 2009;5:e1000300. [PubMed]
- Berget SM. Exon recognition in vertebrate splicing. J Biol Chem 1995;270:2411-4. [PubMed]
- Lev-Maor G, Goren A, Sela N, et al. The “alternative” choice of constitutive exons throughout evolution. PLoS Genet 2007;3:e203. [PubMed]
- Corvelo A, Eyras E. Exon creation and establishment in human genes. Genome Biol 2008;9:R141. [PubMed]
- Kim DS, Huh JW, Kim YH, et al. Bioinformatic analysis of TE-spliced new exons within human, mouse and zebrafish genomes. Genomics 2010;96:266-71. [PubMed]
- Alekseyenko AV, Kim N, Lee CJ. Global analysis of exon creation versus loss and the role of alternative splicing in 17 vertebrate genomes. RNA 2007;13:661-70. [PubMed]
- Sela N, Kim E, Ast G. The role of transposable elements in the evolution of non-mammalian vertebrates and invertebrates. Genome Biol 2010;11:R59. [PubMed]
- Modrek B, Lee CJ. Alternative splicing in the human, mouse and rat genomes is associated with an increased frequency of exon creation and/or loss. Nat Genet 2003;34:177-80. [PubMed]
- Piriyapongsa J, Rutledge MT, Patel S, et al. Evaluating the protein coding potential of exonized transposable element sequences. Biol Direct 2007;2:31. [PubMed]
- Sakharkar MK, Chow VT, Kangueane P. Distributions of exons and introns in the human genome. In Silico Biol 2004;4:387-93. [PubMed]
- Carlton JM, Adams JH, Silva JC, et al. Comparative genomics of the neglected human malaria parasite Plasmodium vivax. Nature 2008;455:757-63. [PubMed]
- Singh N, Sharma R, George A, et al. Cloning and Characterization of Buffalo NANOG Gene: Alternative Transcription Start Sites, Splicing, and Polyadenylation in Embryonic Stem Cell-Like Cells. DNA Cell Biol 2012;31:721-31. [PubMed]
- Carter G, Patel R, Apostolatos A, et al. Protein kinase C delta (PKCδ) splice variant modulates senescence via hTERT in adipose-derived stem cells. Stem Cell Investigation 2014;1:3.
- Fujimoto K, Matsuura K, Hu-Wang E, et al. Thyroid Hormone Activates Protein Arginine Methyltransferase 1 Expression by Directly Inducing c-Myc Transcription during Xenopus Intestinal Stem Cell Development. J Biol Chem 2012;287:10039-50. [PubMed]
- Lee KL, Lim SK, Orlov YL, et al. Graded Nodal/Activin Signaling Titrates Conversion of Quantitative Phospho-Smad2 Levels into Qualitative Embryonic Stem Cell Fate Decisions. PLoS Genet 2011;7:e1002130. [PubMed]
- Mathieu J, Ruohola-Baker H. Regulation of Stem Cell Populations by microRNAs. Adv Exp Med Biol 2013;786:329-51. [PubMed]
- Skreka K, Schafferer S, Nat I-R, et al. Identification of differentially expressed non-coding RNAs in embryonic stem cell neural differentiation. Nucleic Acids Res 2012;40:6001-15. [PubMed]
- Wang C, Wilson-Berryb L, Schedlb T, et al. TEG-1 CD2BP2 regulates stem cell proliferation and sex determination in the C. elegans germ line and physically interacts with the UAF-1 U2AF65 splicing factor. Dev Dyn 2012;241:505-21. [PubMed]
- Gladyshev EA, Arkhipova IR. A widespread class of reverse transcriptase-related cellular genes. Proc Natl Acad Sci U S A 2011;108:20311-6. [PubMed]
- Dhillon B, Cavaletto JR, Wood KV, et al. Accidental amplification and inactivation of a methyltransferase gene eliminates cytosine methylation in Mycosphaerella graminicola. Genetics 2010;186:67-77. [PubMed]
- Braumann I, van den Berg MA, Kempken F. Strain-specific retrotransposon-mediated recombination in commercially used Aspergillus niger strain. Mol Genet Genomics 2008;280:319-25. [PubMed]
- Martinez D, Berka RM, Henrissat B, et al. Genome sequencing and analysis of the biomass-degrading fungus Trichoderma reesei (syn. Hypocrea jecorina). Nat Biotechnol 2008;26:553-60. [PubMed]
- Kämper J, Kahmann R, Bölker M, et al. Insights from the genome of the biotrophic fungal plant pathogen Ustilago maydis. Nature 2006;444:97-101. [PubMed]
- Medstrand P, van de Lagemaat LN, Mager DL. Retroelement distributions in the human genome: variations associated with age and proximity to genes. Genome Res 2002;12:1483-95. [PubMed]
- Shu Y, Li Y, Bai X, et al. Identification and characterization of a new member of the SINE Au retroposon family (GmAu1) in the soybean, Glycine max (L.) Merr., genome and its potential application. Plant Cell Rep 2011;30:2207-13. [PubMed]
- Mourier T, Willerslev E. Does selection against transcriptional interference shape retroelement-free regions in mammalian genomes? PLoS One 2008;3:e3760. [PubMed]
- Baller JA, Gao J, Stamenova R, et al. A nucleosomal surface defines an integration hotspot for the Saccharomyces cerevisiae Ty1 retrotransposon. Genome Res 2012;22:704-13. [PubMed]
- Mularoni L, Zhou Y, Bowen T, et al. Retrotransposon Ty1 integration targets specifically positioned asymmetric nucleosomal DNA segments in tRNA hotspots. Genome Res 2012;22:693-703. [PubMed]
- Novick PA, Smith JD, Floumanhaft M, et al. The evolution and diversity of DNA transposons in the genome of the Lizard Anolis carolinensis. Genome Biol Evol 2011;3:1-14. [PubMed]
- Hawkins JS, Proulx SR, Rapp RA, et al. Rapid DNA loss as a counterbalance to genome expansion through retrotransposon proliferation in plants. Proc Natl Acad Sci U S A 2009;106:17811-6. [PubMed]
- Devos KM, Brown JK, Bennetzen JL. Genome size reduction through illegitimate recombination counteracts genome expansion in Arabidopsis. Genome Res 2002;12:1075-9. [PubMed]
- Loomis WF, Shaulsky G, Wang N. Histidine kinases in signal transduction pathways of eukaryotes. J Cell Sci 1997;110:1141-5. [PubMed]
- Alex LA, Korch C, Selitrennikoff CP, et al. COS1, a two-component histidine kinase that is involved in hyphal development in the opportunistic pathogen Candida albicans. Proc Natl Acad Sci U S A 1998;95:7069-73. [PubMed]
- Larson EM, Idnurm A. Two origins for the gene encoding alpha-isopropylmalate synthase in fungi. PLoS One 2010;5:e11605. [PubMed]
- Zhang Q, Edwards SV. The evolution of intron size in amniotes: a role for powered flight? Genome Biol Evol 2012;4:1033-43. [PubMed]
- Wang D, Su Y, Wang X, et al. Transposon-derived and satellite-derived repetitive sequences play distinct functional roles in Mammalian intron size expansion. Evol Bioinform Online 2012;8:301-19. [PubMed]
- Moss SP, Joyce DA, Humphries S, et al. Comparative analysis of teleost genome sequences reveals an ancient intron size expansion in the zebrafish lineage. Genome Biol Evol 2011;3:1187-96. [PubMed]
- Ohmori Y, Abiko M, Horibata A, et al. A transposon, Ping, is integrated into intron 4 of the DROOPING LEAF gene of rice, weakly reducing its expression and causing a mild drooping leaf phenotype. Plant Cell Physiol 2008;49:1176-84. [PubMed]
- Bergemann M, Lespinet O, M’Barek SB, et al. Genome-wide analysis of the Fusarium oxysporum mimp family of MITEs and mobilization of both native and de novo created mimps. J Mol Evol 2008;67:631-42. [PubMed]
- Zhang XH, Chasin LA. Comparison of multiple vertebrate genomes reveals the birth and evolution of human exons. Proc Natl Acad Sci U S A 2006;103:13427-32. [PubMed]
- Parmley JL, Hurst LD. Exonic splicing regulatory elements skew synonymous codon usage near intron-exon boundaries in mammals. Mol Biol Evol 2007;24:1600-3. [PubMed]
- Spingola M, Grate L, Haussler D, et al. Genome-wide bioinformatic and molecular analysis of introns in Saccharomyces cerevisiae. RNA 1999;5:221-34. [PubMed]
Cite this article as: Zhou K, Kuo A, Grigoriev IV. Reverse transcriptase and intron number evolution. Stem Cell Investig 2014;1:17.


