Philadelphia chromosome-negative acute myeloid leukemia with 11q23/MLL translocation in a patient with chronic myelogenous leukemia
Introduction
Chronic myelogenous leukemia (CML) is defined at molecular level by BCL-ABL1 fusion gene generated from a translocation between chromosome 9q34 and 22q11.2, forming Philadelphia chromosome (Ph) (1). BCR-ABL1 is the only genetic abnormality in 90% of CML cases in chronic phase. As disease progresses, clonal evolution with additional chromosomal changes (ACAs) emerges (2). Approximately 30% of cases in accelerated phase and 50-80% of cases in blast phase have other chromosomal changes besides t(9;22). ACAs often occur in Ph-positive cells (ACAs/Ph+) and are associated with resistance to tyrosine kinase inhibitor (TKI) treatment and disease progression. The most common ACAs include trisomy 8, extra copy of Ph chromosome, i(17)(q10), and trisomy 19. These are so-called major route changes. Other less common ACAs belong to minor route changes.
Occasionally ACAs occur in Ph-negative cells (ACAs/Ph-). Jabbour et al. analyzed cytogenetics in 258 CML patients who were in chronic phase and treated with imatinib (3). After a median follow-up of 37 months, 21 (9%) patients developed chromosomal abnormalities in Ph-negative cells, with −Y and trisomy 8 being the most common. When −Y was excluded, the occurrence rate of ACAs/Ph- was 5%. Although similar cytogenetic changes are associated with myelodysplastic syndromes (MDS) and/or acute myeloid leukemia (AML) in other patients, the emergence of these cytogenetic changes in CML was often transient and disappeared in all but three patients after a median follow-up of 5 months. In a review study by Loriaux et al. (4), the authors summarized 73 CML patients who developed ACAs/Ph- during imatinib treatment. Trisomy 8 was the most common abnormality (53%), followed by chromosome 7 abnormalities (23%). Similar chromosomal changes, also less frequently, have been reported in CML patients treated with interferon (4). Although relatively rare, development of MDS or AML in Ph-negative cells has been reported. Kovitz et al. studied 1,701 CML patients who were treated with imatinib and found that three patients developed AML or MDS in Ph-negative cells (5). Other similar cases have also been reported (6-15). Overall, −7 and complex cytogenetic changes were the most common chromosomal abnormalities in reported cases of MDS and AML that developed in Ph-negative cells. Occasional cases were diploid (5).
In a recent study of the role of clonal evolution with 11q23/MLL rearrangements in CML (16), we identified an interesting case of AML that emerged from a Ph-negative clone in a patient with a history of CML. No similar case of 11q23/Ph-negative AML has been reported in the literature. Thus in this report, we described this case in detail and discussed the potential mechanisms of the emergence of chromosomal abnormalities in Ph-negative cells.
Case presentation
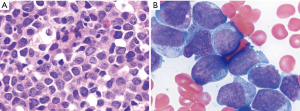
The patient was a 48-year-old man, who initially presented with splenomegaly and leukocytosis with a white blood cell count of 474 K/µL in August, 2010. He was diagnosed with CML, chronic phase. Cytogenetics showed t(9;22)(q34;q11.2)[20]. Molecular study was positive for BCR-ABL1 transcript by real-time reverse transcription polymerase chain reaction (RT-PCR). The patient was treated with several TKIs including nilotinib, imatinib, dasatinib and ponatinib. He responded but experienced intolerance, such as allergy and myelosuppression. In April 2013, the patient presented with pancytopenia with a hemoglobin level of 7 g/dL, white blood cell count of 1.9 K/µL, and platelet count of 10 K/µL. A bone marrow procedure was performed and showed AML with 36% blasts on smears. Flow cytometric immunophenotyping analysis showed blasts were positive for CD13, CD33, CD34 and CD117. Conventional cytogenetics demonstrated the following karyotype: 46,XY,t(11;17)(q23;q25)[20]. The 11q23/MLL gene rearrangement was confirmed by fluorescence in situ hybridization (FISH) analysis. FISH for BCR-ABL1 fusion was detected in 4% interphase nuclei, in keeping with residual CML. The patient was referred to our hospital in May 2013. A repeat bone marrow biopsy showed AML with monocytic differentiation with 63% blasts. Morphologically, blasts were large with moderate amounts of basophilic cytoplasm, round to irregular nuclei, dispersed chromatin and distinct nucleoli (Figure 1). Conventional cytogenetics showed: 46,XY, t(11,17)(q23;q25)[20]. FISH study using MLL dual color, break-apart probe (Abott Molecular, Inc.) showed 94% of cells were positive for MLL gene rearrangement. FISH for BCR/ABL1 on bone marrow was negative at this time. BCR-ABL1/ABL ratio by real-time RT-PCR was 0.09, consistent with the presence of residual CML disease in background. The blasts had the following immunophenotype by flow cytometry: positive for CD4, CD13, CD15, CD33, CD34 partial, CD38, CD64 subset, CD117, CD123, myeloperoxidase, and negative for CD2, CD7, CD14, CD19, CD22, CD36, CD56, HLA-DR, and TdT. The patient was put on clofarabine, idarubicin, and cytarabine along with ponatinib. He showed a transient response with blast percentage falling to 12% on day 21 of therapy, but progressed to 52% blasts on day 28 bone marrow. The patient died 4 months after emergence of 11q23 clone due to disease progression.
Discussion
In this report, we described a unique case of AML developing in Ph-negative cells with 11q23/MLL rearrangement. The summary of its clinicopathological presentation is shown in Table 1. To our best knowledge, this is the first case of 11q23/MLL rearrangement developing in Ph-negative cells in a CML patient. Although discussed in the literature, the exact mechanisms that mediate the development of chromosomal abnormalities in Ph-negative cells are not fully understood. One potential explanation is that TKIs may induce chromosomal changes in Ph-negative cells by inhibiting cellular ABL kinase (c-ABL) activity. C-ABL plays an important role in DNA damage repair following DNA damage (17). The inhibition of c-ABL activity may confer cells to be susceptible to DNA damage signals and induce chromosomal changes. Another possibility is that the pathogenesis of CML is a multiple-step event and t(9;22) is not the earliest event, but rather develops from a Ph-negative hematopoietic stem cell clone. The Ph-negative clone has genetic abnormality that not only induces t(9;22) but also other chromosomal changes. When t(9;22) emerges, the growth advantage of CML cells masks cells with other chromosomal changes. During targeted therapy of CML, Ph-positive cells are eliminated and cells with ACAs/Ph- re-emerge.

Full table
Rearrangements of 11q23/MLL are common and present in 70-80% of cases of infant acute leukemia (18). In adult, 11q23/MLL rearrangements are less frequent and seen in de novo and therapy-related AML (19). The presence of 11q23/MLL in CML is very rare (16). The translocation partners for 11q23/MLL are diverse with t(9;11)(p22;q23), t( 4;11)(q21;q23), and t(11;19)(q23;p13) being the most common (20). T(11;17)(q23;q25) involving SEPT9 gene on 17q25 described in our case has been reported in AML and MDS with a poor prognosis (21). Consistently, the patient reported here showed poor response to chemotherapy and died 4 months after the diagnosis of AML.
Acknowledgements
None.
Footnote
Conflicts of Interest: The authors have no conflicts of interest to declare.
Written informed consent was obtained from the patient for publication of this case report and accompanying images.
References
- Rowley JD. Letter: A new consistent chromosomal abnormality in chronic myelogenous leukaemia identified by quinacrine fluorescence and Giemsa staining. Nature 1973;243:290-3. [PubMed]
- Baccarani M, Deininger MW, Rosti G, et al. European LeukemiaNet recommendations for the management of chronic myeloid leukemia: 2013. Blood 2013;122:872-84. [PubMed]
- Jabbour E, Kantarjian HM, Abruzzo LV, et al. Chromosomal abnormalities in Philadelphia chromosome negative metaphases appearing during imatinib mesylate therapy in patients with newly diagnosed chronic myeloid leukemia in chronic phase. Blood 2007;110:2991-5. [PubMed]
- Loriaux M, Deininger M. Clonal cytogenetic abnormalities in Philadelphia chromosome negative cells in chronic myeloid leukemia patients treated with imatinib. Leuk Lymphoma 2004;45:2197-203. [PubMed]
- Kovitz C, Kantarjian H, Garcia-Manero G, et al. Myelodysplastic syndromes and acute leukemia developing after imatinib mesylate therapy for chronic myeloid leukemia. Blood 2006;108:2811-3. [PubMed]
- Georgiou G, Efthymiou A, Vardounioti I, et al. Development of acute myeloid leukemia with NPM1 mutation, in Ph-negative clone, during treatment of CML with imatinib. Leukemia 2012;26:824-6. [PubMed]
- Jin Huh H, Won Huh J, Myong Seong C, et al. Acute lymphoblastic leukemia without the Philadelphia chromosome occurring in chronic myelogenous leukemia with the Philadelphia chromosome. Am J Hematol 2003;74:218-20. [PubMed]
- Perel JM, McCarthy C, Walker O, et al. Clinical significance of development of Philadelphia-chromosome negative clones in patients with chronic myeloid leukemia treated with imatinib mesylate. Haematologica 2005;90 Suppl:ECR25. [PubMed]
- Quintás-Cardama A, Kantarjian H, Abruzzo LV, et al. Extramedullary BCR-ABL1-negative myeloid leukemia in a patient with chronic myeloid leukemia and synchronous cytogenetic abnormalities in Philadelphia-positive and -negative clones during imatinib therapy. Leukemia 2007;21:2394-6. [PubMed]
- Chee YL, Vickers MA, Stevenson D, et al. Fatal myelodysplastic syndrome developing during therapy with imatinib mesylate and characterised by the emergence of complex Philadelphia negative clones. Leukemia 2003;17:634-5. [PubMed]
- Cherrier-De Wilde S, Rack K, et al. Philadelphia-negative acute lymphoblastic leukemia developing in a CML patient in imatinib mesylate-induced complete cytogenetic remission. Leukemia 2003;17:2046-8. [PubMed]
- Skalska-Sadowska J, Januszkiewicz-Lewandowska D, Derwich K, et al. Ph-negative isolated myeloid sarcoma with NPM1 gene mutation in adolescent with Ph-positive chronic myeloid leukemia in remission after treatment with allogeneic bone marrow transplantation and imatinib mesylate. Pediatr Blood Cancer 2015;62:1070-1. [PubMed]
- Schafhausen P, Dierlamm J, Bokemeyer C, et al. Development of AML with t(8;21)(q22;q22) and RUNX1-RUNX1T1 fusion following Philadelphia-negative clonal evolution during treatment of CML with Imatinib. Cancer Genet Cytogenet 2009;189:63-7. [PubMed]
- Dvorak P, Hruba M, Subrt I. Development of acute myeloid leukemia associated with Ph-negative clone with inv(3)(q21q26) during imatinib therapy for chronic myeloid leukemia. Leuk Res 2009;33:860-1. [PubMed]
- Wakim JJ, Tirado CA, Dowell J, et al. The first case of Philadelphia chromosome-negative acute promyelocytic leukemia following imatinib for chronic myelogenous leukemia. Cancer Genet 2012;205:124-7. [PubMed]
- Wang W, Tang G, Cortes JE, et al. Chromosomal rearrangement involving 11q23 locus in chronic myelogenous leukemia: a rare phenomenon frequently associated with disease progression and poor prognosis. J Hematol Oncol 2015;8:32. [PubMed]
- Wang X, Zeng L, Wang J, et al. A positive role for c-Abl in Atm and Atr activation in DNA damage response. Cell Death Differ 2011;18:5-15. [PubMed]
- Tomizawa D, Koh K, Sato T, et al. Outcome of risk-based therapy for infant acute lymphoblastic leukemia with or without an MLL gene rearrangement, with emphasis on late effects: a final report of two consecutive studies, MLL96 and MLL98, of the Japan Infant Leukemia Study Group. Leukemia 2007;21:2258-63. [PubMed]
- Schoch C, Schnittger S, Klaus M, et al. AML with 11q23/MLL abnormalities as defined by the WHO classification: incidence, partner chromosomes, FAB subtype, age distribution, and prognostic impact in an unselected series of 1897 cytogenetically analyzed AML cases. Blood 2003;102:2395-402. [PubMed]
- Meyer C, Hofmann J, Burmeister T, et al. The MLL recombinome of acute leukemias in 2013. Leukemia 2013;27:2165-76. [PubMed]
- Lee SG, Park TS, Oh SH, et al. De novo acute myeloid leukemia associated with t(11;17)(q23;q25) and MLL-SEPT9 rearrangement in an elderly patient: a case study and review of the literature. Acta Haematol 2011;126:195-8. [PubMed]
Cite this article as: Jaitly V, Wang W, Hu S. Philadelphia chromosome-negative acute myeloid leukemia with 11q23/MLL translocation in a patient with chronic myelogenous leukemia. Stem Cell Investig 2015;2:13.


